Plant Biology
Bridging the Volumetric Gap in Plant Ultrastructure
For decades, the standard approach to investigating plant cell ultrastructure relied upon two-dimensional transmission electron microscopy (2D TEM) of ultrathin resin sections. While TEM offers extraordinary nanometer-level resolution, extracting reliable three-dimensional (3D) information from 2D micrographs has historically required laborious manual serial sectioning, a technique highly susceptible to section loss, distortion, and significant alignment errors [14, 29, 254]. More recently, electron tomography has provided excellent 3D resolution, but it is fundamentally limited to exceedingly small observation volumes, rendering whole-cell or whole-organelle reconstructions practically impossible [228, 246, 247].
Serial Block-Face Scanning Electron Microscopy (SBF-SEM) has emerged as a transformative technology that bridges the critical gap between the wide field of view provided by fluorescence light microscopy and the ultra-high resolution of electron tomography [3, 17, 33, 37]. By integrating an automated ultramicrotome directly within the high-vacuum specimen chamber of a field-emission scanning electron microscope, SBF-SEM continuously alternates between slicing a resin-embedded tissue block (typically removing 15–100 nm layers) and imaging the newly exposed block face using backscattered electrons [13, 14, 17, 20, 32, 34, 37, 200]. This highly automated "slice-and-view" methodology generates perfectly registered, massive stacks of TEM-like images, enabling the 3D reconstruction of large tissue volumes—up to hundreds of cubic micrometers—at nanoscale resolution [5, 8, 18, 40, 107]. While initially developed for neurobiology and connectomics [10, 11, 17, 37, 251], SBF-SEM is now being rapidly adopted in plant biology, revolutionizing our spatial understanding of cell wall architecture, complex organelle morphology, and intercellular communication networks [37, 97, 252].
Overcoming Plant-Specific Preparation Challenges
Plant tissues present a highly unique and challenging microenvironment for volume electron microscopy. Unlike animal tissues, mature plant cells feature an osmotically induced intracellular hydrostatic (turgor) pressure that can approach or exceed 1 MPa, which places immense elastic strain on the surrounding extracellular matrix [48, 117]. Furthermore, plant cells are encased in rigid, complex cellulosic cell walls and often protected by waxy cuticles that severely impede the diffusion of chemical fixatives, heavy metal stains, and embedding resins [52, 103, 159]. Consequently, direct application of standard mammalian SBF-SEM preparation protocols (such as the traditional osmium-thiocarbohydrazide-osmium, or OTO, methods) frequently produces severe structural artifacts in plant specimens, including collapsed and undulating cell walls or highly distorted, star-shaped vacuoles [44, 48].
To mitigate these issues, specialized modifications to tissue processing have been developed. For resin infiltration, low-viscosity Spurr's epoxy resin is widely preferred over other media because it penetrates plant cell walls effectively and possesses the requisite hardness to withstand both the physical forces of in-chamber diamond knife sectioning and the thermal stress of repeated electron beam scanning [8]. Because SBF-SEM detects backscattered electrons from the block face, post-sectioning staining on grids is impossible; therefore, the entire tissue block must be rendered highly electron-dense and conductive en bloc [8, 12]. To achieve this, plant biologists have resurrected and adapted various selective contrasting techniques. The zinc iodide-osmium (ZIO) impregnation method has proven exceptional for selectively staining plant endomembrane systems, clearly contrasting the nuclear envelope, endoplasmic reticulum (ER), Golgi apparatus, and plastid thylakoids [37]. Alternatively, the application of uranyl acetate followed by a combination of copper and lead citrate provides outstanding contrast for polysaccharides, making it highly suitable for visualising cell walls and Golgi-derived secretory vesicles [37].
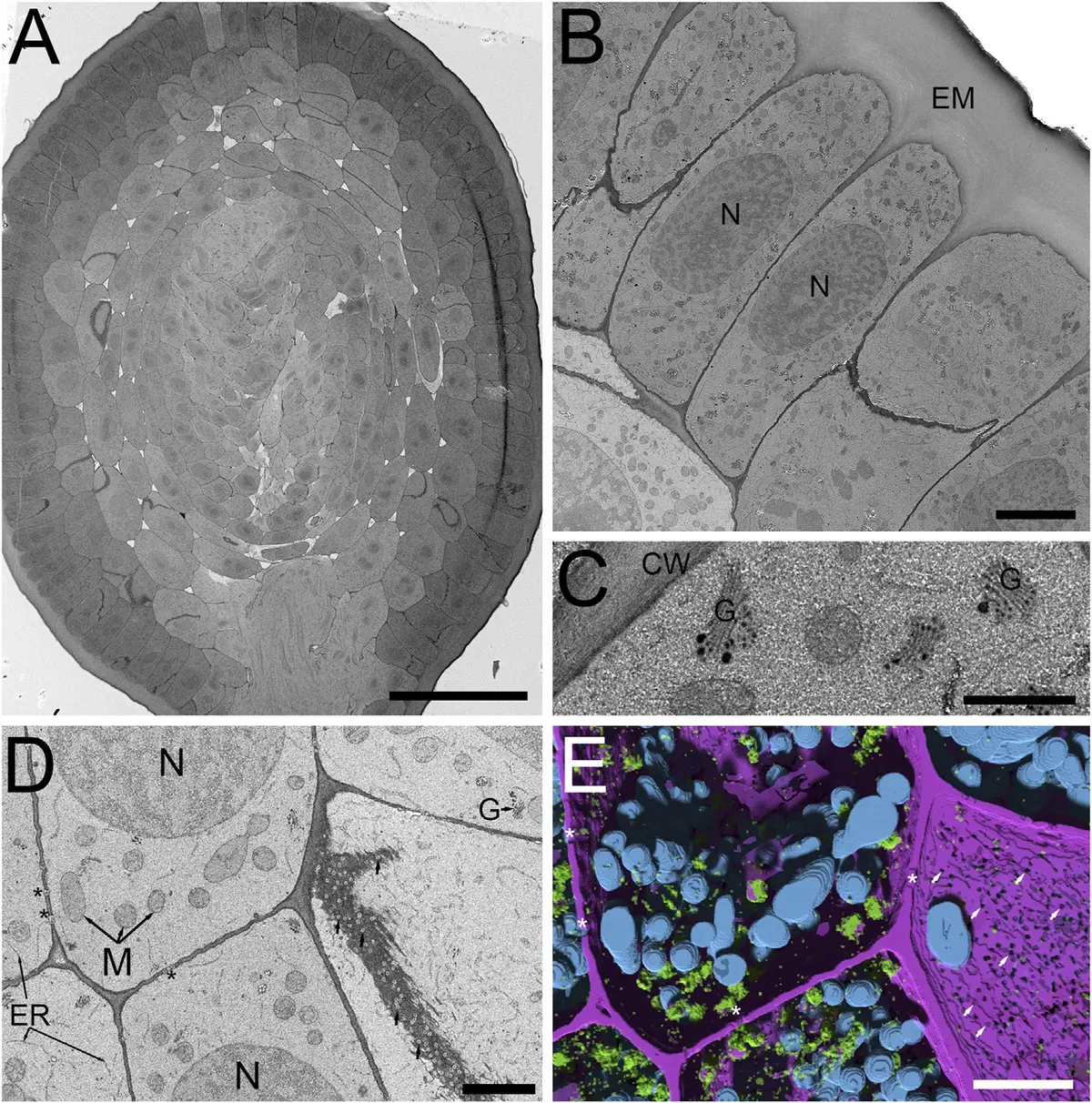
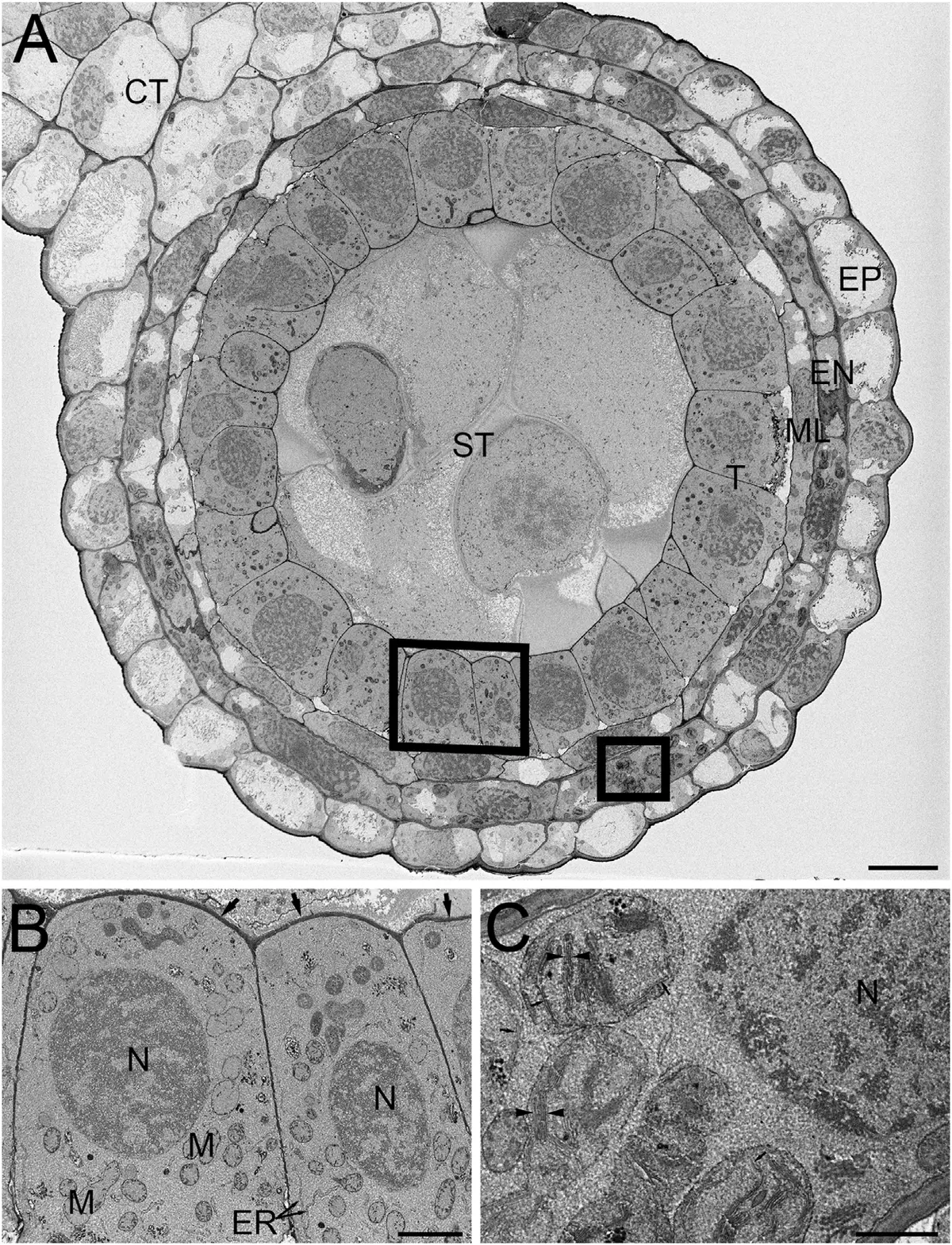
Advanced cryofixation techniques have also been adapted for SBF-SEM. High-pressure freezing followed by freeze-substitution (HPF-FS) circumvents the artifacts induced by slow chemical cross-linking [44, 228]. A novel aqueous freeze-substitution protocol (FSaqOTO) has successfully preserved the high-density ground cytoplasmic matrix of *Hordeum vulgare* (barley) root apical meristems and anthers [44]. This protocol drastically enhances the electron density of plant cell walls and vacuolar compartments, facilitating subsequent machine-learning-based automated segmentation [44] (Figures 49–50).
Chloroplast Architecture and Biogenesis
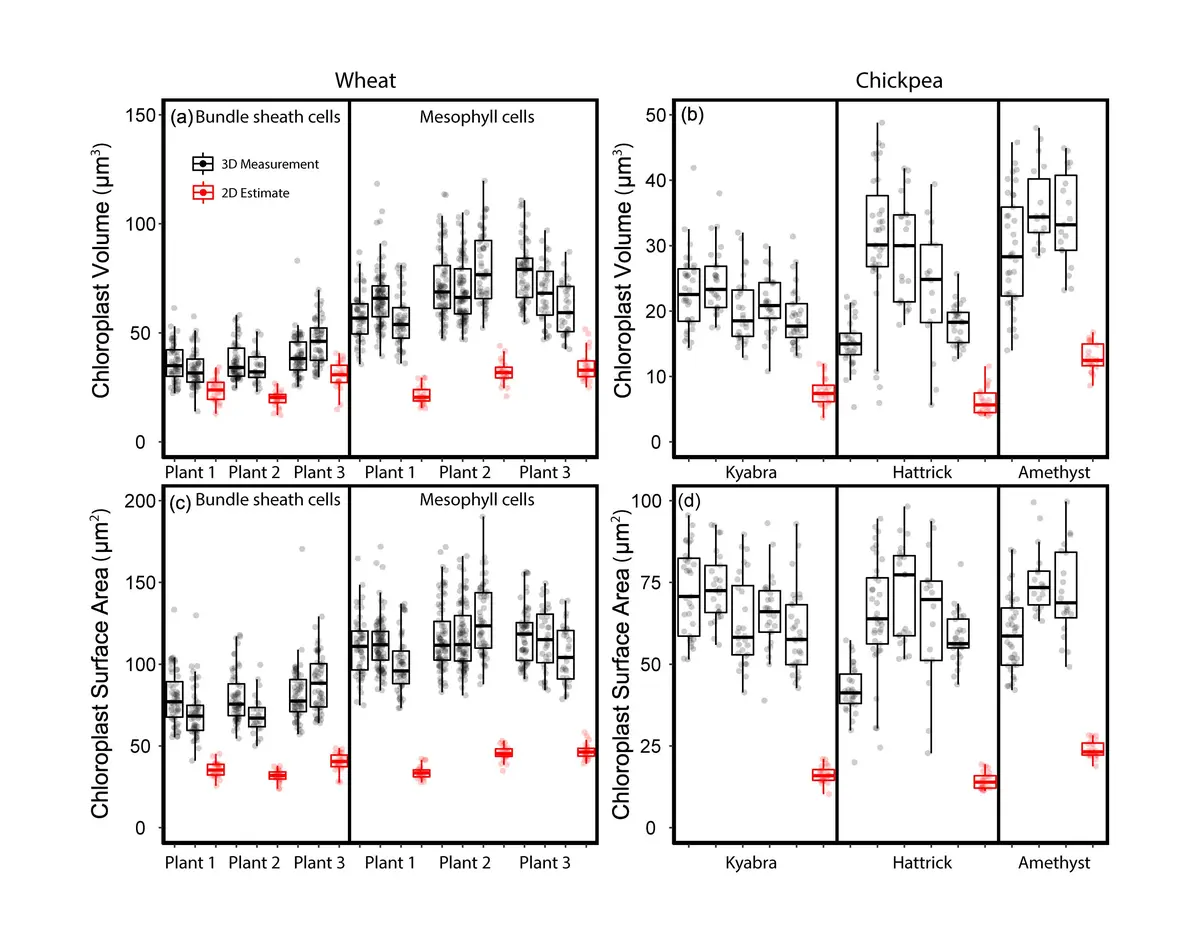
One of the most profound impacts of SBF-SEM in plant biology has been the correction of systemic volumetric biases in chloroplast morphometrics. Historically, organelle volumes were extrapolated from 2D TEM or light microscopy images using idealized geometric models, such as spheres or triaxial ellipsoids [83, 129, 152, 228]. However, plastids frequently possess irregular, flattened, or concavely lobed morphologies that defy simple geometry [83]. SBF-SEM studies of wheat and chickpea leaves have definitively proven that 2D TEM extrapolation systematically underestimates the true volume of mesophyll chloroplasts by up to 60-61%, and bundle sheath chloroplasts by 45% [83]. Similarly, simple geometric equations miscalculate the surface-area-to-volume ratios of complex, lobed spongy mesophyll cells [83].
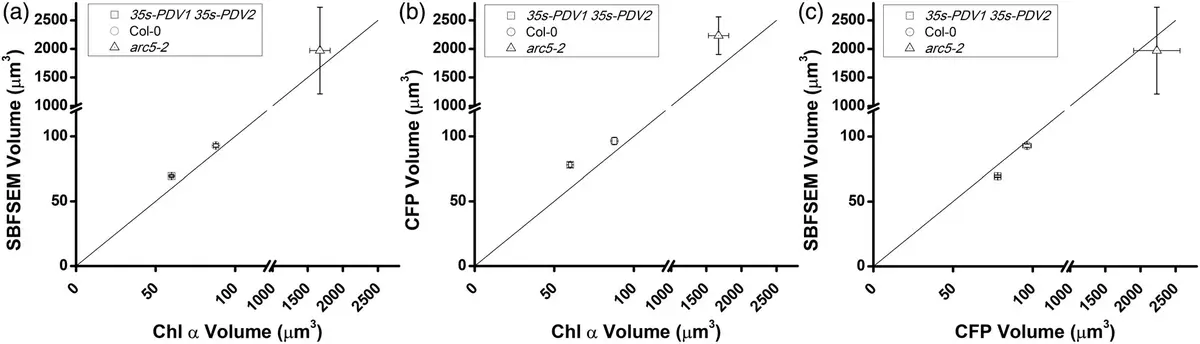
While confocal laser scanning microscopy (LSM) allows for *in situ* live imaging without the shrinkage artifacts of chemical fixation (spinach chloroplasts, for example, can shrink by 30% during standard TEM fixation), LSM suffers from limited z-axial resolution and fluorescence scattering [84, 152]. Furthermore, LSM cannot clearly resolve the internal stroma from the thylakoid membranes, and software often erroneously merges tightly packed chloroplasts into single objects [84, 129, 152]. SBF-SEM overcomes these optical limitations, providing unambiguous, high-resolution 3D segmentations of individual plastids across large tissue expanses [84, 152]. In C4 crop species such as maize and sugarcane, SBF-SEM has quantified dramatic dimorphism in chloroplast architecture, demonstrating that bundle sheath chloroplasts are two- to three-fold larger than their mesophyll counterparts and occupy a staggering 60–70% of the total bundle sheath cell volume [129] (Figures 51–52).
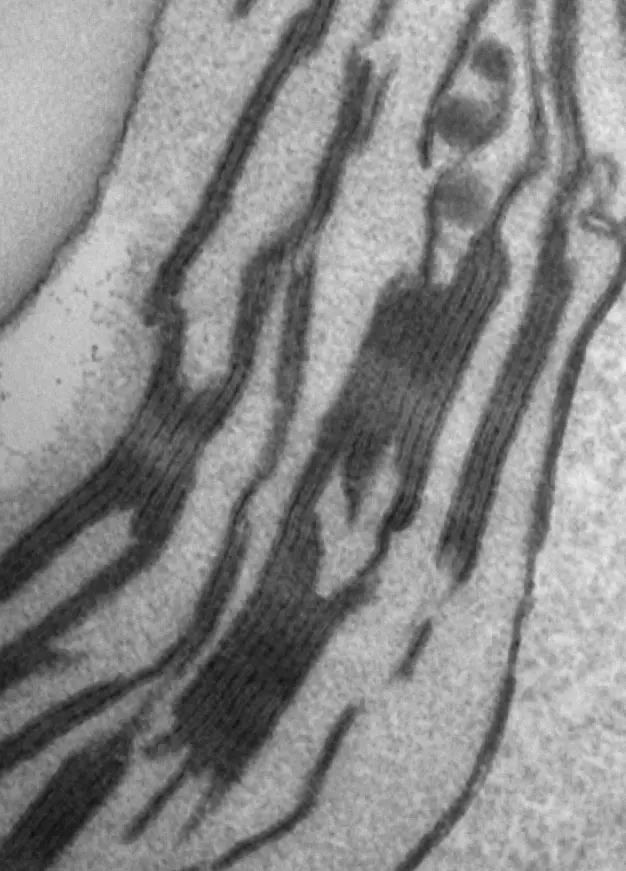
Beyond steady-state anatomy, SBF-SEM has provided unprecedented, quantitative insights into chloroplast biogenesis. During the de-etiolation (greening) of *Arabidopsis thaliana* cotyledons, SBF-SEM tracked the ultrastructural transformation of etioplasts into functional chloroplasts over a 96-hour continuous light exposure period [246, 247]. Early in this process (T0 to T4 hours), 3D reconstructions captured the rapid disassembly of paracrystalline prolamellar bodies (PLBs)—which initially consist of 0.8–1 μm hexagonal units and tubular prothylakoids—into elongated, planar lamellar sheets [246, 247]. By 24 to 96 hours, these structures differentiate into distinct appressed grana stacks and non-appressed stroma lamellae [246, 247]. SBF-SEM enabled the exact calculation of this membrane proliferation: by 96 hours, the total thylakoid surface area within the seedling cotyledons reaches approximately 700 mm², an extraordinary 500-fold expansion relative to the external leaf surface area [246, 247]. Integrating these morphometric models with lipidomic profiles of galactolipids (MGDG and DGDG) and immunodetection of photosynthetic complexes revealed that chloroplast biogenesis occurs in two distinct, previously unappreciated phases—an initial massive structural establishment phase followed by a subsequent chloroplast division phase [228, 246] (Figure 53).
Plasmodesmata and Intercellular Communication
Understanding symplastic transport requires accurate mapping of plasmodesmata—nanoscale, plasma membrane-lined channels that bridge the thick cellulosic cell walls of adjacent plant cells. While individual plasmodesmata tubules are exceedingly narrow (< 25 nm) and can occasionally be lost between the 40–50 nm sectioning intervals of standard SBF-SEM [129], the technology is highly proficient at mapping entire plasmodesmata pit fields in 3D [129]. In C4 leaves, SBF-SEM spatial analyses revealed that these pit fields occupy roughly 2–3% of the total cell surface area at both the mesophyll-mesophyll and bundle sheath-mesophyll interfaces [129].
In *Arabidopsis thaliana*, SBF-SEM has been deployed to quantitatively map plasmodesmata distribution within critical phloem unloading zones, specifically at the interfaces between sieve elements, phloem pole pericycle cells, and the endodermis [228]. Generating an accurate topological map of these connections is essential for informing computational models of phloem fluxes and whole-plant resource allocation [228]. Interestingly, under specific freeze-substitution sample preparation protocols (like FSaqOTO), plasmodesmata present with an inverted contrast—appearing electron-transparent against a highly electron-dense cell wall—which greatly facilitates their automated detection in 3D datasets [44].
SBF-SEM has also brought to light more dramatic forms of intercellular communication. A striking example is intercellular nuclear migration (INM) in the male meiocytes of plants. Long debated and frequently dismissed as a mechanical artifact of squashed slide preparations, INM is nearly impossible to study dynamically because meiocytes are buried deep within layers of protective anther tissue [251, 252]. By utilizing SBF-SEM to reconstruct entire, intact *Nicotiana tabacum* (tobacco) and rye anther locules, researchers provided definitive 3D proof of massive INM events during the first meiotic prophase [108, 251]. SBF-SEM datasets revealed that 90–100% of normal tobacco meiocytes participate as either nuclear donors or recipients, sharing genetic material through expansive cytomictic channels without the formation of micronuclei [251, 252]. The migrating nuclei actively establish direct physical contacts with the native nuclei of recipient cells, a phenomenon entirely hidden from standard 2D TEM [228, 251, 252].
Root Tip Development, Endomembranes, and Pathogen Interactions
The dense, highly dynamic cytoplasm of meristematic cells represents a prime target for volumetric reconstruction. SBF-SEM has been widely utilized to map the topological complexity of the plant endomembrane system in developing root tips of Arabidopsis, pea, and maize [83]. By selectively staining membranes with the ZIO technique, researchers have reconstructed continuous networks of cortical and tubular ER transitioning into cisternal ER sheets, juxtaposed with independent Golgi stacks [31, 33, 48]. Crucially, SBF-SEM resolved a long-standing optical illusion derived from 2D sections: what often appears as dozens of fragmented, distinct vacuoles in a single thin section of a meristematic root cell is revealed in 3D to be a single, highly continuous, convoluted organellar network [31, 48].
The technology has similarly resolved spatial relationships in specialized storage tissues. In the developing maize endosperm, 2D and confocal microscopy left ambiguities regarding the exact biogenesis of protein bodies (PBs) [234]. SBF-SEM screening of large tissue volumes demonstrated unequivocally that zein-containing PBs bud directly from central areas of continuous ER membrane sheets, maintaining direct connections within the native ER network rather than forming as isolated vesicles [228, 234]. Similar broad reorganizations have been mapped in the digestive glands of the Venus flytrap (*Dionaea muscipula*), where SBF-SEM captured the complex remodeling of the secretory endomembrane system following mechanical prey stimulation [228].
Volumetric imaging is also advancing the field of plant pathology. In studies of Arabidopsis cotyledons infected by the hemibiotrophic fungal pathogen *Colletotrichum higginsianum*, SBF-SEM illuminated the 3D topology of host-pathogen interfaces [250]. Reconstructions showcased primary intracellular biotrophic hyphae winding extensively through host epidermal cells [250]. The large field of view enabled researchers to track the pathogen's induction of massive host membrane disintegration, including the engulfment of host cytoplasm by the central vacuole and the striking proliferation of interconnected, tubular ER bodies wrapping around the invasive hyphae [250]. Furthermore, nano-resolution volume EM has mapped the intricate 3D feeding interactions between sap-sucking psyllid insects (*Trioza camphorae*) and *Cinnamomum camphora* tissues [253].
Wood, Xylem, and Complex Tissue Anatomy
The rigid, highly structured nature of plant vascular elements presents a perfect use-case for SBF-SEM. Beyond the delicate structures of the mesophyll and roots, the technique excels at evaluating the massive secondary cell walls of lignified tissues. For instance, SBF-SEM was utilized to evaluate the progressive 3D cell wall reinforcement and lignification of vein cells during the maturation of hydrangea decorative sepals [248]. By assembling stacks of backscattered electron images acquired at 100-nm intervals, researchers constructed highly accurate topological models of the xylem and surrounding supportive tissues, correlating structural thickness directly to physical desiccation resistance [248]. The technique is comparably powerful for investigating hierarchical porosities and highly ordered matrices in other robust biological materials, mirroring its use in characterizing resilient mammalian structures [97].
Image Analysis, Deep Learning, and the Future
While the hardware of modern SBF-SEM systems permits the automated, uninterrupted acquisition of thousands of registered images over a matter of days [17, 30], the subsequent data extraction remains an immense bottleneck [17, 28, 35, 228]. Extracting morphometric data requires the segmentation of structures—outlining boundaries of chloroplasts, mitochondria, or cell walls across hundreds of sequential slices [96, 201, 247, 249]. Manual segmentation is phenomenally time-consuming, often taking thousands of hours for a single large dataset, severely limiting high-throughput comparative biology [17, 35, 228].
To overcome this, the field is rapidly integrating artificial intelligence and machine learning pipelines. Advanced algorithms based on 3D convolutional neural networks and flood-filling networks are being trained specifically on plant ultrastructure [48, 108, 117]. Software packages such as Sensor 3D [44] and the PyTorch Connectomics-based Anatomics MLT [48, 84] have demonstrated remarkable accuracy in autonomously labeling chloroplast volumes, mitochondria, Golgi stacks, and cell walls. By marrying these sophisticated AI segmentation tools with enhanced en bloc staining protocols that minimize physiological artifacts [48], SBF-SEM is evolving from a purely qualitative, descriptive tool into a robust, high-throughput quantitative platform.
In conclusion, Serial Block-Face Scanning Electron Microscopy has fundamentally shifted the paradigm of plant ultrastructural analysis. By illuminating the true 3D architecture of chloroplast networks, mapping precise symplastic and endomembrane continuities, and exposing the hidden dynamics of cell-to-cell nuclear migration, SBF-SEM provides an indispensable structural foundation. As automated segmentation tools continue to mature, the data generated will increasingly power the next generation of physiological and metabolic computational models across the plant sciences.