Cardiac and Muscle Biology
Methodological Advances in Muscle Volume Electron Microscopy
In the realm of cardiac and muscle biology, serial block-face scanning electron microscopy (SBF-SEM) has emerged as a transformative imaging modality that bridges the spatial divide between the expansive fields of view offered by confocal fluorescence microscopy and the ultra-high, yet volume-restricted, resolution of electron tomography [14, 24, 100]. By automating the sequential acquisition of backscattered electron images from the freshly sectioned surface of a resin-embedded tissue block, SBF-SEM permits the three-dimensional (3D) reconstruction of entire cells and tissue microenvironments [13, 24]. The ability to capture massive tissue volumes—often exceeding tens of thousands of cubic micrometres—at intermediate nanometre-scale resolutions (typically 10–15 nm in the X-Y plane and 30–70 nm in the Z-direction) has facilitated unprecedented quantitative analyses of muscle ultrastructure in both health and disease [24, 42, 100, 223, 225] (Figures 44–48).
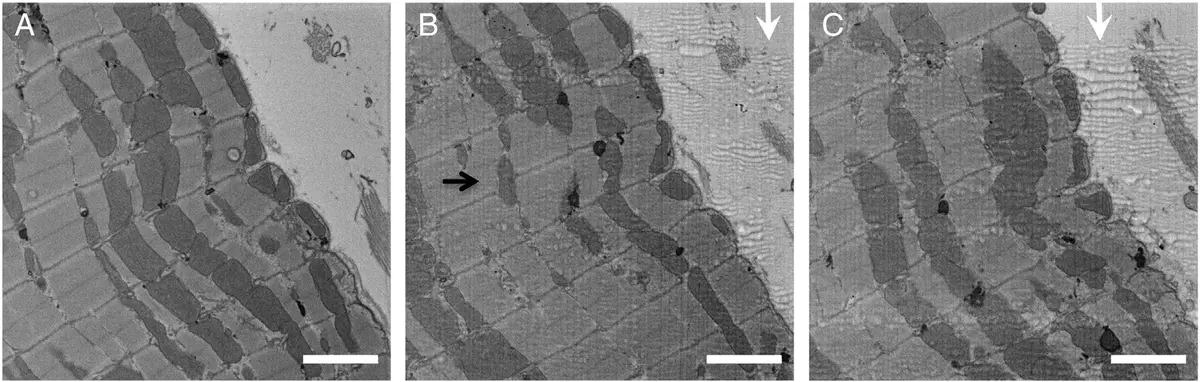

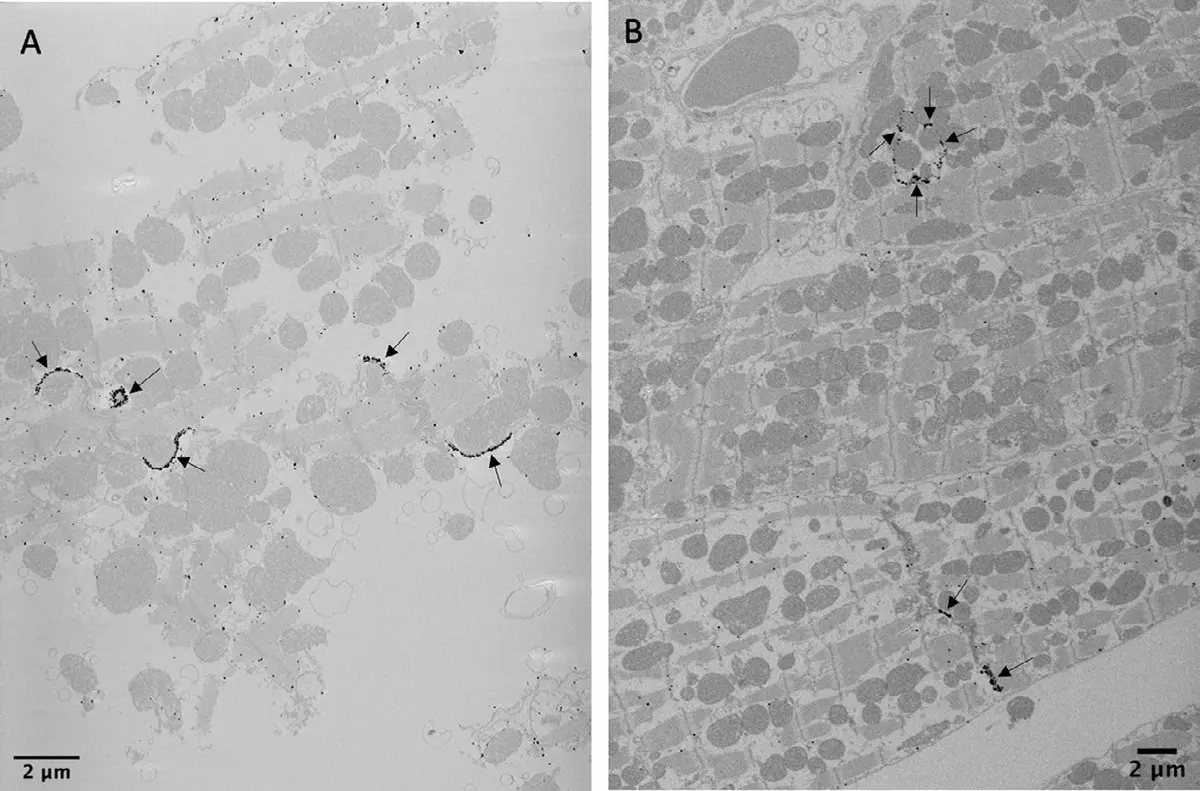
Achieving high-fidelity SBF-SEM datasets of muscle tissue requires rigorous, specialised sample preparation protocols designed to maximize electron density and mitigate charging artefacts [8, 26, 77, 107]. Muscle tissues, such as the left ventricular myocardium, are generally fixed using Karnovsky’s fixative—a tightly controlled cocktail of paraformaldehyde and glutaraldehyde in sodium cacodylate buffer—to preserve native cellular morphology without inducing osmotic shrinkage [24, 100, 223, 240]. To provide the necessary contrast for backscattered electron detection, specimens are subjected to en bloc heavy metal staining, frequently utilising a modified OTO (osmium-thiocarbohydrazide-osmium) protocol followed by uranyl acetate and lead aspartate [12, 116, 223, 240]. For specific subcellular targeting, such as the sarcoplasmic reticulum (SR), reduced osmium protocols incorporating potassium ferrocyanide or high concentrations of calcium chloride can be employed to render the target membranes intensely electron-dense, appearing virtually black against the lighter myofilaments [24]. Stained tissues are subsequently dehydrated and embedded in hard plastic resins, such as Durcupan ACM, which provides the structural stability required for the continuous, automated slicing by the ultramicrotome's diamond knife within the high-vacuum or low-vacuum environment of the scanning electron microscope [34, 100, 223, 240]. Prior to serial sectioning, techniques like micro-computed tomography (microCT) are sometimes utilised to verify the spatial orientation of muscle fibres within the resin block, ensuring that the acquired block-face images align optimally with the longitudinal or transverse axes of the tissue [100].
3D Architecture of the Healthy Cardiomyocyte
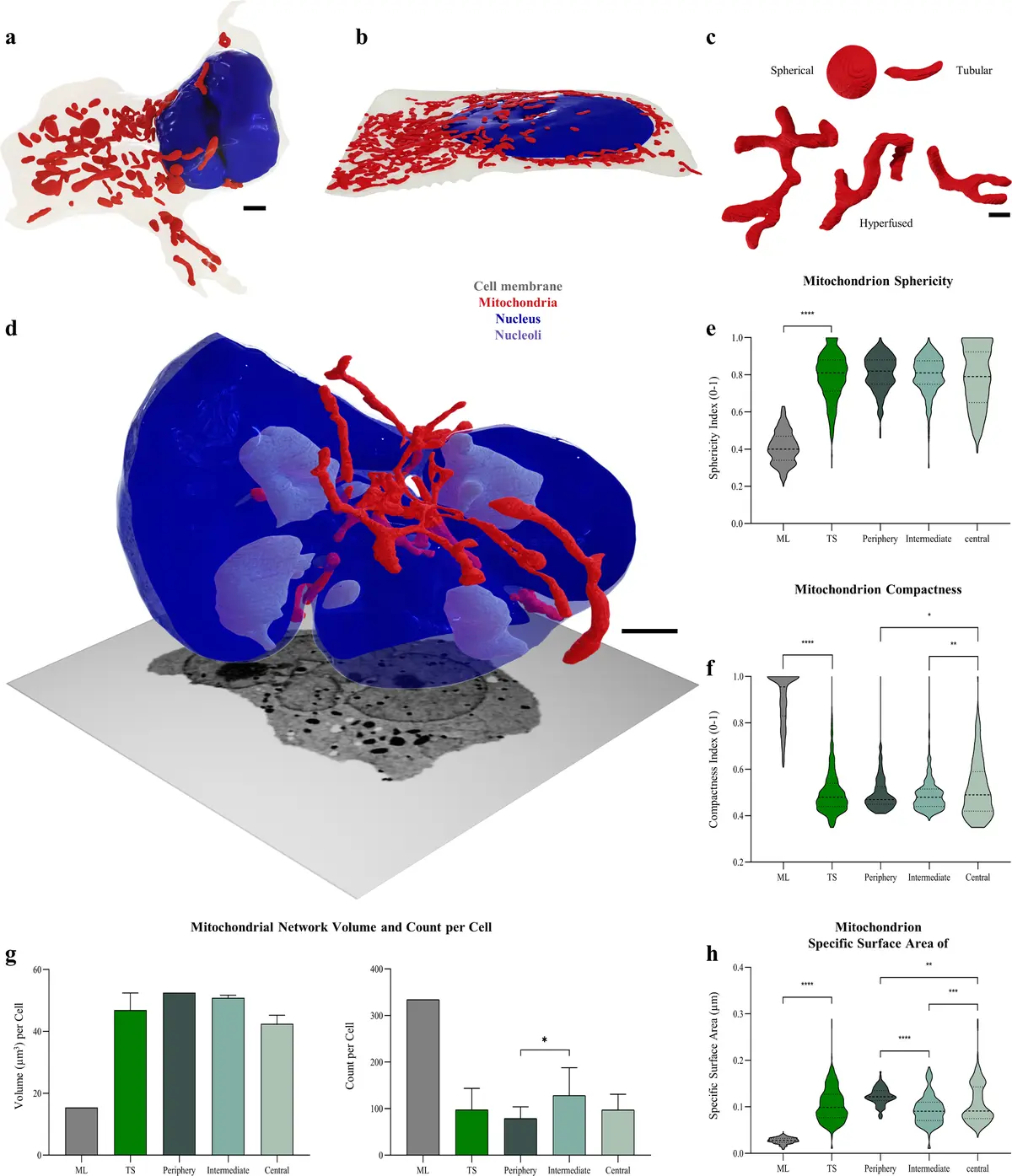
The application of automated and semi-automated segmentation algorithms to large-volume SBF-SEM datasets has allowed for the complete 3D reconstruction of isolated, intact adult cardiac myocytes within their native myocardial syncytium [100, 225]. Detailed volumetric analyses of single rat ventricular cardiomyocytes reveal that the internal space is densely packed and highly compartmentalised, with the contractile apparatus (myofibrils) occupying approximately 46–48% of the cell volume, while the energetic machinery (mitochondria) constitutes a remarkably high 50–53% [100]. In contrast, the nuclear volume accounts for less than 1% of the total intracellular space [100]. SBF-SEM segmentations also highlight the complex structural topology of the cardiomyocyte, mapping the branching trajectories of myofibrils as they connect with adjacent cells, and detailing the heterogeneous distribution of organelles, such as the increased clustering of mitochondria within the subsarcolemmal space, at cellular branching points, and in the perinuclear region [100].
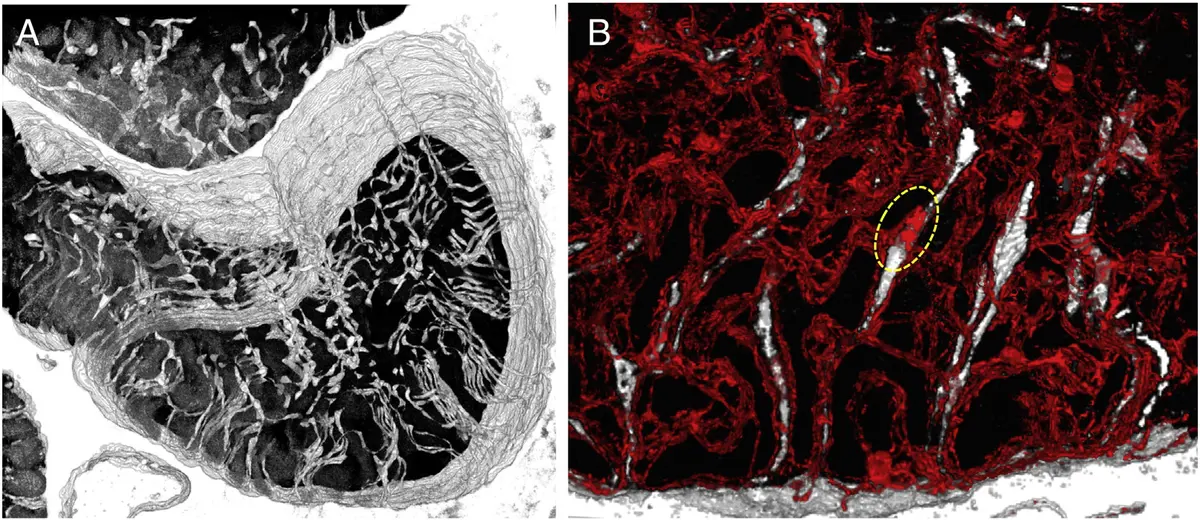
#### The Sarcotubular Network and Dyad Organization A critical domain of investigation utilising SBF-SEM in cardiac biology is the 3D mapping of the sarcotubular network, comprising the transverse tubules (t-tubules) and the SR [14, 24]. T-tubules are deep invaginations of the sarcolemma that facilitate the rapid, uniform propagation of action potentials into the cell interior [24]. In 3D renderings generated from transverse sections of healthy sheep myocardium, the t-tubule network exhibits a highly organized, spoke-like radial geometry, extending centripetally from the sarcolemma towards the cell centre [24, 238]. In longitudinal views, these tubules align periodically along the sarcomeric Z-lines, weaving perpendicular to the sarcolemma [238]. SBF-SEM has established that mammalian ventricular t-tubules possess an average diameter ranging between 200 and 400 nm [24, 238], and typically penetrate just under half the width of the myocyte [238]. High-resolution block-face imaging has resolved previously unappreciated morphological nuances, including the presence of dilations or "nodules" occurring along the length and at the tips of the invaginations [24], as well as the occasional presentation of "twin tubules"—pairs of t-tubules traversing the cell side-by-side at a single Z-line [238].
The intimate spatial relationship between the t-tubules and the junctional SR (jSR) forms the dyad, the fundamental structural unit of cardiac excitation-contraction coupling [24]. By employing selective, electron-dense SR staining procedures, researchers can easily threshold SBF-SEM image stacks to isolate and segment the SR [24]. These segmentations demonstrate how the SR network weaves intricately through the myofilaments and drapes along the edges of the mitochondria to connect individual dyads [24]. Volumetric interrogation of these datasets allows for the precise calculation of jSR volume and surface area, parameters that are critical for computational biophysicists seeking to estimate the accommodation and clustering patterns of ryanodine receptors (RyRs) within the calcium microdomain [24, 225].
#### Mitochondrial Subpopulations and Networking SBF-SEM has proven invaluable for characterising the morphological diversity and spatial networking of mitochondria within the working myocardium. In healthy cardiac tissue, mitochondria are classically categorised into distinct subpopulations based on their anatomical locale: subsarcolemmal mitochondria (SSM) situated beneath the plasma membrane, interfibrillar mitochondria (IFM) arrayed in parallel rows between the myofibrils, and perinuclear mitochondria (PNM) grouped adjacent to the nuclear envelope [125, 223, 243]. Shape classification analyses of SBF-SEM datasets reveal that these subpopulations possess distinct morphometric profiles; IFM are predominantly oval and arranged in elongated strands, whereas SSM and PNM are frequently irregular, dynamically molding their shapes to match the undulating topography of the sarcolemma or the nucleus [243]. Beyond basic geometry, SBF-SEM's continuous Z-axis resolution captures highly complex networking behaviours, including the presence of trans-mitochondrial "nanotunnels"—slender, tubular extensions that physically bridge adjacent mitochondria across the sarcomeric Z-lines, facilitating long-distance electrical and biochemical communication [237, 241].
#### Intercalated Discs and Cardiac Cellular Junctions The application of SBF-SEM has also been extended to the study of intercellular connections within the heart. Correlative methodologies combining nanogold immunolabelling with silver enhancement have enabled the specific subcellular visualisation of gap junction proteins, such as Connexin 43 (Cx43), within adult murine hearts [127]. In lower vertebrates, SBF-SEM has been deployed to map the complex junctional region (JR) separating the spongy and compact layers of the zebrafish ventricle, revealing an intricate, three-dimensional network of resident fibroblasts and phenotypically transitional cardiomyocytes connected by adherens junctions [240].
Ultrastructural Remodeling in Cardiac Pathology
The ability of SBF-SEM to assess hundreds of cubic micrometres of tissue at a nanometre resolution has made it an indispensable tool for uncovering the ultrastructural basis of cardiac disease [24, 100]. In large animal models of myocardial infarction (MI), SBF-SEM has unmasked profound, region-dependent remodeling of the t-tubular network [238]. While remote regions of the myocardium may retain a relatively normal architecture, cardiomyocytes located within the infarct border zone exhibit severe t-tubule disorganization, manifesting as a loss of transverse alignment with the Z-lines and widespread tubule fragmentation [238]. Notably, volume EM reveals that these fragmented remnants, which are completely disconnected from the extracellular space, still possess an inner basement membrane, confirming their sarcolemmal origin [238]. This fragmentation is accompanied by a significant overall reduction in t-tubule density within the border zone, contributing directly to the blunted calcium transients and contractile dysfunction characteristic of post-MI pathology [238].
In the context of heart failure (HF), SBF-SEM reconstructions of both animal and human samples indicate extensive structural disarray [100, 237]. In human dilated cardiomyopathy, intermyofibrillar mitochondria exhibit tremendous inter-sample variability and display significant increases in volume, surface area, and perimeter compared to healthy controls [237]. The failing mitochondria frequently deviate from normal spherical or oval morphologies, adopting highly complex, pathological phenotypes including "donut" shapes and excessive nanotunnel branching [237]. Concurrently, the myofibrillar apparatus in failing human hearts demonstrates a loss of circularity, an increased cross-sectional area, and a higher frequency of aberrant myofibril branching [237]. Intriguingly, in models studying recovery from HF, atrial myocytes display a unique pattern of t-tubule remodeling wherein highly disordered t-tubules weave erratically between enlarged, irregularly arranged mitochondrial beds, fundamentally altering the spatial proximity of calcium release sites to the myofilaments [239].
SBF-SEM is equally powerful in exploring metabolic disorders and ischaemic injuries. In the early stages of diabetic cardiomyopathy, quantitative 3D shape analyses show a differential remodeling of mitochondrial subpopulations, marked by a specific reduction in the ratio of oval to irregular mitochondria within the interfibrillar compartment [243]. Furthermore, in models of cardiac ischaemia-reperfusion injury, SBF-SEM is utilised to assess the internal integrity of mitochondria, enabling investigators to visually distinguish and quantify populations of functionally connected, networked cristae versus pathologically fragmented or "orphaned" cristae [224].
Skeletal Muscle Ultrastructure, Aging, and Disease
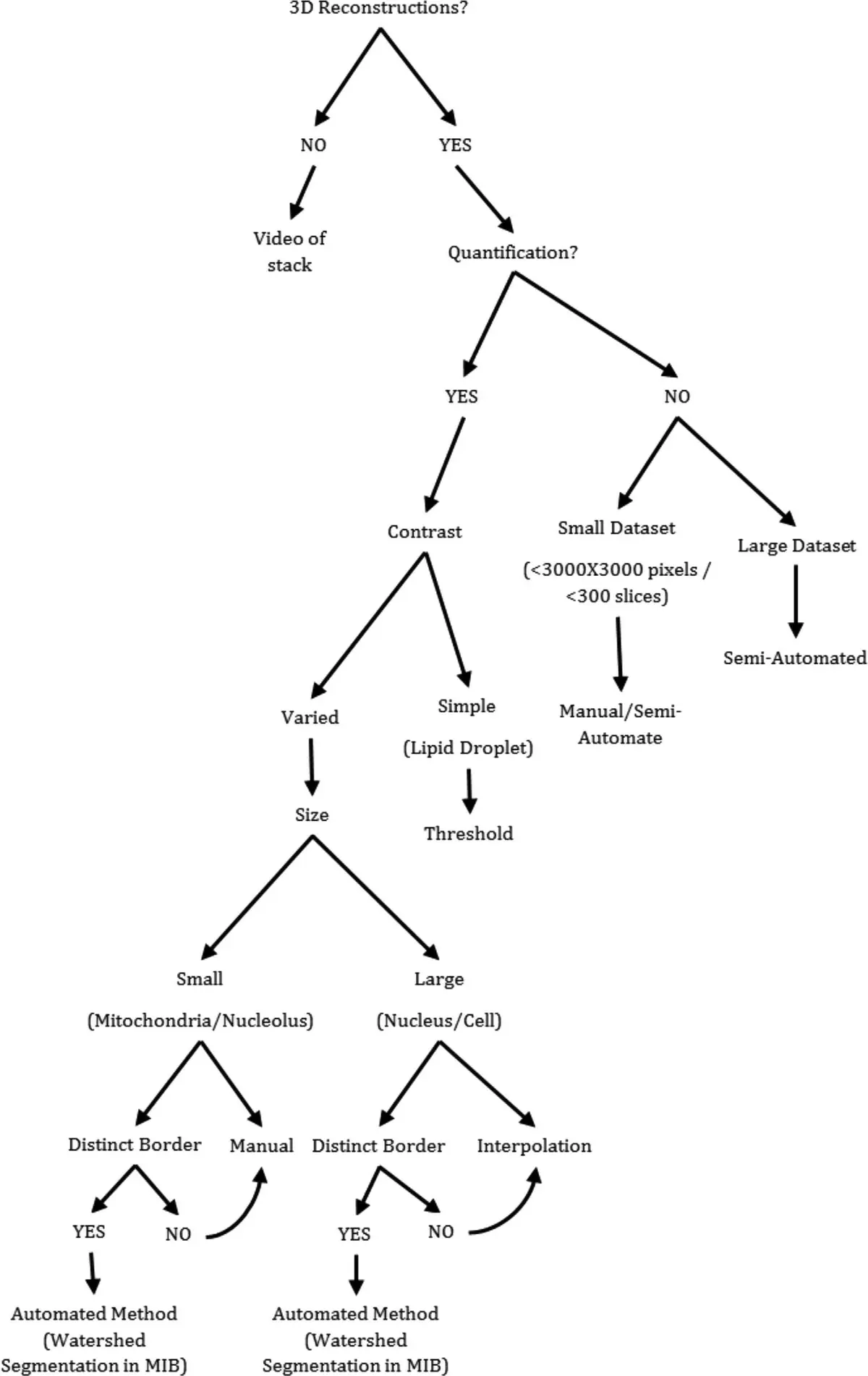
The utility of SBF-SEM extends broadly across skeletal muscle biology, shedding light on developmental morphogenesis, fibre type specialisation, and the morphological hallmarks of ageing [35, 236]. Image analysis platforms—such as Fiji (with TrakEM2), Amira, and Microscopy Image Browser (MIB)—are routinely employed to segment complex SBF-SEM datasets of foetal and adult skeletal muscle, utilizing a combination of manual contour tracing, contrast-based thresholding, and interpolation to reconstruct myofibrils, multiple mitochondrial populations, Z-lines, and lipid droplets [35].
#### Fibre Type Diversity and Age-Related Remodeling Skeletal muscle exhibits immense heterogeneity in its metabolic profile, which is reflected at the ultrastructural level. Comparative SBF-SEM studies of distinct muscle groups, such as the highly oxidative broiler gastrocnemius (predominantly type I slow-twitch fibres) versus the glycolytic pectoralis major (predominantly type IIb fast-twitch fibres), reveal stark differences in mitochondrial topology [241]. Type I fibres feature a dense, highly interconnected 3D mitochondrial network characterised by extensive trans-mitochondrial nanotunnels bridging individual organelles, alongside frequent interactions with lipid droplets; conversely, type IIb fibres present a much sparser, highly fragmented spatial network [241].
During the ageing process, skeletal muscle undergoes progressive functional decline (sarcopenia) intimately linked to mitochondrial deterioration [112, 236]. SBF-SEM volumetric renderings of muscles from adolescent versus aged murine cohorts (e.g., gastrocnemius and soleus) demonstrate that ageing provokes a marked reduction in mitochondrial size, total volume, surface area, and perimeter [236]. Notably, while the mitochondrial networks in skeletal muscle become severely fragmented and highly heterogeneous with age, the mitochondrial populations within aged cardiac tissue remain comparatively homogeneous [236].
#### Molecular Drivers of Mitochondrial Morphology The high-throughput nature of SBF-SEM has accelerated research into the molecular machinery governing mitochondrial dynamics, notably the mitochondrial contact site and cristae organizing system (MICOS) [236]. Targeted ablation of MICOS complex subunits, such as Sam50, in both murine and human skeletal myotubes leads to dramatic alterations in 3D architecture [227]. Volumetric segmentation reveals that the loss of Sam50 precipitates profound mitochondrial fragmentation, drastically reducing mitochondrial volume, surface area, and perimeter, while disrupting overall network complexity [227].
#### Correlative Approaches in Human Mitochondrial Myopathy In clinical contexts, SBF-SEM has been successfully integrated with functional enzymatic assays to investigate human mitochondrial myopathies [125, 242]. A pioneering correlative technique known as COX-SBFSEM couples traditional Cytochrome c Oxidase (COX) histochemistry with serial block-face imaging to map the per-mitochondrion oxidative phosphorylation status in situ [125, 244]. In muscle biopsies from patients harbouring large-scale mitochondrial DNA (mtDNA) deletions, this multimodal approach can distinguish between COX-positive (functional), COX-intermediate, and COX-deficient mitochondria within a single muscle fibre [125, 244]. 3D reconstructions reveal that while COX-positive mitochondria maintain robust, large, and complex geometries, mitochondria within COX-deficient fibres are invariably spherical, undersized, and highly fragmented, providing a direct visual link between metabolic dysfunction and structural collapse [125, 244].
Smooth Muscle and the Extracellular Matrix
Beyond the intracellular space of striated muscle, SBF-SEM has significantly advanced the understanding of smooth muscle and the structural organisation of the musculoskeletal extracellular matrix (ECM) [116, 245]. Because the ECM is notoriously difficult to image due to a lack of intrinsic electron density, researchers have adapted contrast-enhancing procedures, such as ruthenium red staining, to visualise extensive perimysial collagen structures over hundreds of micrometres [245]. This capability is critical for understanding the mechanical basis of muscle fibrosis and stiffness [245]. Similarly, in the study of embryonic tendon development, SBF-SEM has been utilised to track individual collagen fibrils across large volumes, revealing that fibrils undergo complex intertwining rather than remaining strictly columnar as they mature [6, 12]. In the cardiovascular system, SBF-SEM datasets generated from the mouse aorta have elucidated the nanoscopic arrangement of fibrillin micro-fibrils within concentric elastin sheets, offering a geometrically accurate, high-resolution 3D view of the elastic fibre systems that regulate arterial compliance [116].
Collectively, the integration of SBF-SEM into muscle and cardiac biology has catalysed a transition from two-dimensional observations to comprehensive, three-dimensional spatial understanding. By allowing the detailed segmentation and volumetric quantification of myofibrils, sarcotubular networks, mitochondria, and extracellular matrices across unprecedented spatial scales, volume electron microscopy continues to uncover the intricate structure-function relationships governing muscle physiology in health, ageing, and disease.