Future Directions and Challenges
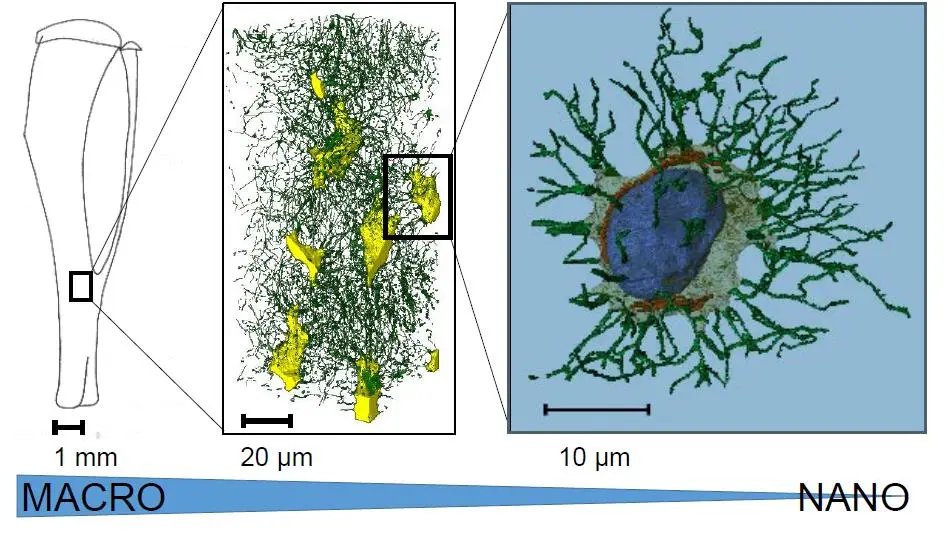
Serial Block-Face Scanning Electron Microscopy (SBF-SEM) has emerged as a cornerstone of modern volume electron microscopy (vEM), driving a "quiet revolution" in our ability to visualize the three-dimensional ultrastructure of cells, tissues, and organisms [9], [9], [168]. By utilizing an ultramicrotome mounted directly inside the vacuum chamber of a scanning electron microscope (SEM), SBF-SEM bypasses the labor-intensive and error-prone manual collection of serial sections required by traditional transmission electron microscopy (TEM) [12], [71], [14]. However, the technique remains intrinsically destructive; the sample is milled away during acquisition, meaning sections cannot be revisited [14], [22], [14], [18], [18]. Furthermore, as researchers push the boundaries of SBF-SEM to examine larger volumes at higher resolutions, the technique faces several persistent bottlenecks [16]. The future of SBF-SEM lies in a multidisciplinary approach: optimizing sample preparation to mitigate charging, engineering faster and more sensitive detectors, integrating correlative multimodal imaging, and harnessing artificial intelligence (AI) to manage the unprecedented deluge of data (Figure 67).
Overcoming Sample Preparation and Charging Bottlenecks
A fundamental limitation of SBF-SEM imaging is the accumulation of negative electron charge on the block face. Biological samples are intrinsically non-conductive, and the insulating plastic resins used for embedding exacerbate charging artifacts [8], [29], [22]. When a specimen charges, it causes image distortion, loss of signal-to-noise ratio (SNR), and physical instability during sectioning, such as skipped cuts or "chatter" from the diamond knife [14], [77], [50]. While optimized heavy metal en bloc staining protocols—such as the reduced osmium-thiocarbohydrazide-osmium (rOTO) method—provide essential backscattered electron (BSE) contrast and introduce some internal conductivity, charging remains a severe problem for lipid-poor tissues, highly porous samples, and cultured cell monolayers [8], [26], [29], [22], [77].
To circumvent these issues, imaging is frequently conducted in variable pressure (low-vacuum) modes by introducing water vapor or nitrogen gas into the SEM chamber to dissipate charge [14], [65], [12], [66], [15]. However, low-vacuum imaging inherently scatters the primary electron beam, slightly degrading lateral resolution and SNR [14], [15]. Consequently, there is a concerted effort to develop novel sample preparation workflows that enable high-vacuum imaging. One promising direction is the development of conductive resins formulated with additives like carbon black or Ketjen Black [18], [8]. While these resins successfully dissipate charge, they render the block visually opaque, complicating macroscopic sample orientation and the localization of embedded structures prior to sectioning [8].
Alternatively, "minimal resin" (MR) embedding techniques have been introduced to drastically reduce the volume of insulating plastic surrounding the tissue [29], [29]. By eliminating the excess empty resin, MR embedding effectively mitigates surface charging and significantly facilitates the navigation to surface-linked regions of interest (ROIs), preventing accidental trimming of the target tissue [29], [29], [29]. Within the SEM chamber itself, technological solutions such as Focal Charge Compensation (FCC)—which utilizes a localized micro-injection of nitrogen gas directly over the block face—have proven highly effective in neutralizing charge without compromising the high-vacuum environment of the broader chamber [18], [11], [5]. Additionally, automated in-chamber sputter coating between cuts has been shown to effectively eliminate charging and increase SNR, though this approach extends the overall acquisition time [18], [14], [18].
Hardware Innovations: Improved Detectors and Faster Acquisition
SBF-SEM relies on the detection of backscattered electrons (BSE) at relatively low primary beam energies (typically 1.0 to 2.5 keV) to limit the electron interaction volume and thereby enhance the axial (z) resolution [14], [19], [25], [77]. Because BSE yield is naturally low at these voltages, detector sensitivity is paramount [19]. Manufacturers have responded by developing highly specialized, low-energy BSE detectors. For instance, the Gatan OnPoint BSE detector has been specifically engineered to offer heightened sensitivity at low accelerating voltages (<5 kV), enabling sub-microsecond pixel dwell times while maintaining high SNR [77]. Similarly, custom low-energy BSE (LE BSE) detectors integrated into TESCAN and ConnectomX Katana systems feature modified geometries that permit imaging at extremely short working distances, further maximizing signal collection [19], [19], [19], [19].
Despite these detector enhancements, SBF-SEM acquisition remains a lengthy process; capturing a large volume often requires days to continuous weeks of uninterrupted imaging [29], [16], [93]. For massive "connectomics" projects aiming to map entire mammalian brain regions, single-beam SEM acquisition speeds are a hard physical bottleneck [71], [16], [14]. To resolve this, the field is moving toward parallelization. Multibeam SEM (mSEM) technology increases throughput by splitting a single electron source into an array of parallel beamlets [16], [14], [16], [9]. Current mSEM architectures utilize 61, 91, or even 196 beams simultaneously to acquire large tile grids in a fraction of the traditional time [159], [38], [16]. A recent commercial manifestation of this concept is FAST-EM, which employs 64 parallel beams combined with a unique optical transmission detection system to rapidly capture ultrastructural data [178], [178], [16]. While currently most mSEM systems are optimized for array tomography (imaging pre-cut serial sections on solid substrates or tapes) [22], [16], [9], [178], the eventual integration of multibeam technology with in-chamber SBF-SEM microtomy holds the potential to reduce data acquisition times from years to a matter of weeks [38], [16].
Pushing the Limits of Axial Resolution
A persistent physical limitation of SBF-SEM is the severe anisotropy of the generated voxels [18], [100], [143]. While the lateral (x, y) resolution of a field-emission SEM can easily reach 3 to 5 nm, the axial (z) resolution is dictated by the minimum slice thickness that the in-chamber ultramicrotome can reliably cut without causing sample damage [32], [13], [14], [14], [32]. For most optimally prepared samples, this minimum cutting thickness is reliably between 25 and 50 nm [32], [13], [50], [58], [38], [32]. While slices as thin as 10 nm have occasionally been achieved under ideal conditions [50], [38], routine isotropic resolution remains the domain of Focused Ion Beam SEM (FIB-SEM), which utilizes a gallium or plasma ion beam to mill sections as thin as 5 nm, albeit over vastly smaller fields of view [32], [38], [5], [32], [9], [18].
To bridge this z-resolution gap in SBF-SEM, researchers have developed innovative computational and multi-energy imaging strategies [18], [18]. One promising approach involves dual primary energy imaging combined with Monte Carlo simulations of electron scattering (using software such as CASINO) [145], [25], [58], [25], [145]. By capturing successive images of the identical block-face at differing accelerating voltages—such as 1.0 keV and 1.4 keV—the electron beam penetrates to distinctly different depths (e.g., approximately 13 nm and 25 nm, respectively) [25], [25]. A linear relationship between the ratio of these backscattered signals and the depth of heavy-metal-stained features allows algorithms to deconvolve the data, effectively reconstructing sub-surface structures and extracting a virtual z-resolution of 5 to 12.5 nm from a 25 nm physical slice [145], [25], [25], [145]. Systems like the ThermoFisher Teneo Volumescope natively integrate this multi-energy optical sectioning with mechanical slicing to vastly improve volumetric resolution [18], [66], [18], [18].
The Growing Importance of CLEM for Targeted Volume EM
As SBF-SEM expands beyond basic structural mapping into functional cell biology, the inability of electron microscopy to inherently identify specific proteins or molecular states becomes a distinct disadvantage [14], [9], [184]. Furthermore, locating a rare or dynamic cellular event—such as a specific mitotic stage or a scarce pathogen interaction—within a massive, visually homogenous tissue block is functionally akin to finding a needle in a haystack [26], [5], [28]. To address this, Correlative Light and Electron Microscopy (CLEM), and specifically volume CLEM (vCLEM), has become an indispensable methodology [46], [14], [3], [46], [9], [14].
In vCLEM workflows, functional or genetic information is first captured using 3D fluorescence light microscopy (LM), such as confocal or two-photon (2P) microscopy [50], [46], [3], [27], [3]. The sample is then fixed, embedded, and subjected to high-resolution SBF-SEM [14], [124], [46], [184]. Combining these datasets allows researchers to achieve "correlation for relocation"—using fluorescence to find the precise ROI to trim and scan—and "correlation for registration"—superimposing the molecular identity over the nanometer-scale ultrastructure [9], [9].
Targeting the precise ROI in a completely opaque resin block requires sophisticated fiducial marking [77], [91], [14]. Researchers have successfully utilized 2P near-infrared branding (NIRB) to burn precise 3D coordinate fiducial marks into the tissue prior to SBF-SEM processing [50], [50], [50]. Once the block is in the SEM chamber, the operator uses the NIRB marks to locate the exact cell of interest, allowing the SBF-SEM acquisition parameters to be "gated"—shrinking the field of view and lowering the slice thickness (e.g., to 80 nm or less) specifically around the target [91]. To further streamline this pipeline and eliminate the physical sample transfer that introduces orientation mismatch, highly advanced integrated microscopes have been developed [38], [14], [9]. Systems that build a miniature fluorescence or reflection light microscope directly into the SBF-SEM microtome stage (such as the "miniLM") allow sequential in-chamber acquisition of both modalities within a shared coordinate space, drastically reducing alignment errors [38], [14], [3].
Computationally registering the LM and SBF-SEM volumes remains a formidable challenge due to extreme disparities in resolution, contrast, and non-linear tissue shrinkage [124], [124], [197]. Historically, alignment required tedious manual landmark placement using tools like BigWarp or ec-CLEM [124]. However, automated registration algorithms are rapidly advancing. Point-cloud-based pipelines such as CLEM-Reg utilize deep learning models (like MitoNet) to automatically segment ubiquitous biological landmarks—such as mitochondria—in both the fluorescence and EM volumes [124], [124], [124]. By applying Laplacian of Gaussian (LoG) filters and dynamic thresholding, CLEM-Reg generates corresponding point clouds that computationally warp and align the multimodal volumes with minimal user intervention [124].
Taming the Data Deluge: Integration with AI and Image Analysis
The unprecedented automation of SBF-SEM has solved the problem of data acquisition, but it has created a massive new bottleneck: data processing and analysis [16], [76], [19], [18], [14], [14], [14]. A single SBF-SEM run can easily encompass thousands of images, generating datasets that range from hundreds of gigabytes to several petabytes [50], [16], [16], [16]. In traditional workflows, human experts must manually trace the membranes of cells, organelles, or neurites slice-by-slice to generate 3D reconstructions [18], [163]. This manual segmentation is staggeringly time-consuming; dense circuit reconstruction in connectomics can require tens of thousands to literally thousands of years of human labor per cubic millimeter of tissue [14].
Consequently, the integration of Artificial Intelligence (AI) and Machine Learning (ML) is no longer optional; it is a critical necessity [16], [10], [28]. Deep Convolutional Neural Networks (CNNs), particularly U-Net architectures, are revolutionizing the automated pixel-level classification and instance segmentation of vEM data [90], [162], [165], [287]. These deep learning models have successfully automated the labeling of complex structures ranging from simple nuclei and mitochondria to highly tortuous neuronal circuits and interdigitated cellular organoids [90], [165], [161], [287].
However, deep learning models are inherently data-hungry, requiring massive amounts of manually annotated "ground truth" data for training [76], [162], [14]. Because SBF-SEM image quality frequently suffers from varying contrast, out-of-focus regions, anisotropic z-resolution, and local charging artifacts, a model trained on one tissue type often fails when applied to another [100], [14], [163]. To bypass the prohibitive cost of generating new training data for every experiment, researchers are increasingly leveraging transfer learning [162], [162], [163]. By taking a pre-trained generalized model—such as Cellpose—and "fine-tuning" it with a remarkably small subset of newly annotated SBF-SEM images (e.g., densely packed platelet masses), segmentation accuracy can be dramatically improved in a fraction of the time [162], [162].
Furthermore, AI is being deployed earlier in the pipeline for image restoration and registration [9], [176]. Self-supervised deep learning methods like Noise2Void or Noise2Noise can dynamically denoise raw SBF-SEM images, enhancing the SNR of fast, low-dose acquisitions before they even reach the segmentation stage [145], [9]. For dataset alignment, optical-flow-based algorithms (such as the Diffeomorphic Demons algorithm) are being adapted to perform non-rigid registration, effectively smoothing out the geometric distortions and physical slice-to-slice jitter that inherently plague SBF-SEM cutting [100], [16], [176].
For researchers without extensive programming expertise, modern software suites are bridging the gap. Programs such as Amira, Microscopy Image Browser (MIB), 3DMOD, and Fiji offer highly integrated, semi-automated segmentation toolkits (including magic wand thresholding, watershed algorithms, and z-axis interpolation) that greatly accelerate the extraction of quantitative metrics from SBF-SEM volumes [19], [35], [81], [18], [81], [35], [35], [35], [35], [35].
Accessibility, Standardization, and Data Sharing
As SBF-SEM matures from a highly specialized technique into a mainstream biological tool, maximizing its accessibility is crucial [12], [9], [12], [9], [14]. Successfully executing a volume EM project requires multidisciplinary expertise spanning specialized heavy-metal tissue fixation, complex ultramicrotomy, vacuum physics, and advanced computational rendering [286], [26], [18]. To democratize access, institutions are increasingly centralizing SBF-SEM hardware within dedicated imaging core facilities [285], [286], [19], [38], [16], [46], [46]. Lightweight, modular ultramicrotomes (such as the ConnectomX Katana) that can be easily mounted and dismounted from a standard SEM stage allow core facilities to utilize a single microscope for both standard high-resolution SEM and SBF-SEM, heavily reducing financial barriers [19], [19], [19], [18], [19], [18].
Software accessibility is equally important. While commercial systems often rely on proprietary control interfaces (e.g., Gatan DigitalMicrograph), the development of open-source Python-based acquisition software like SBEMimage has revolutionized experiment control [9], [19], [74], [74], [74]. SBEMimage communicates directly with the SEM’s low-level application programming interface (API), bypassing scanning speed limitations (capable of up to 40 MHz acquisition rates via SmartSEM) [74], [74], [74]. Crucially, it provides highly flexible control paradigms tailored for week-long runs, including adaptive multi-ROI tile grids, automated debris detection, algorithmic auto-focusing, and remote user monitoring [19], [19], [74], [74].
Finally, as volume EM generates petabytes of rich ultrastructural data, the biological community faces a pressing need for standardized data sharing [16], [16], [10], [16]. SBF-SEM datasets are incredibly dense; a single volume imaged for a specific study (e.g., mapping a neural circuit) contains massive amounts of unanalyzed data regarding mitochondria, endoplasmic reticulum, vasculature, and glia [10], [169]. The establishment of open-access, cloud-based repositories—such as the Electron Microscopy Public Image Archive (EMPIAR), the BioImage Archive, and OpenOrganelle—allows researchers worldwide to download and mine existing 3D datasets, maximizing the scientific return on investment [18], [10], [136], [16], [169]. For this open-science ecosystem to function, the community must universally adopt rigorous reporting standards. Detailed documentation of specimen preparation, resin type, section thickness, electron dose, and the specific segmentation algorithms utilized must be uniformly published alongside SBF-SEM datasets to ensure long-term reproducibility and cross-study comparison [17], [8], [35], [8], [16].
Conclusion
Serial Block-Face Scanning Electron Microscopy has irrevocably transformed our understanding of three-dimensional cellular architecture. Moving forward, the continued evolution of SBF-SEM relies on harmonizing hardware, chemistry, and computer science. By developing smarter conductive resins and deploying ultra-fast multibeam detectors, researchers will overcome the physical limits of charging and acquisition speed. Concurrently, the tight integration of vCLEM workflows and deep learning-based segmentation will unlock the molecular identity and functional context of these vast ultrastructural landscapes. Through centralized core facilities, open-source control software, and robust data-sharing repositories, SBF-SEM is poised to become an accessible, high-throughput standard for the global life sciences community.