AI and Machine Learning for Segmentation
The Quantitative Bottleneck in Volume Electron Microscopy
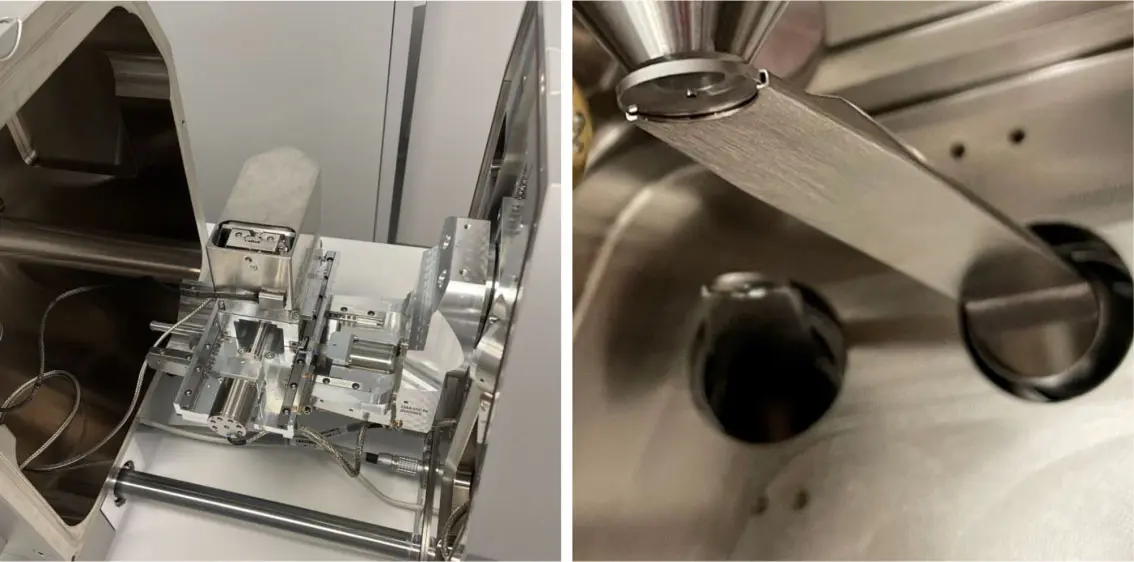
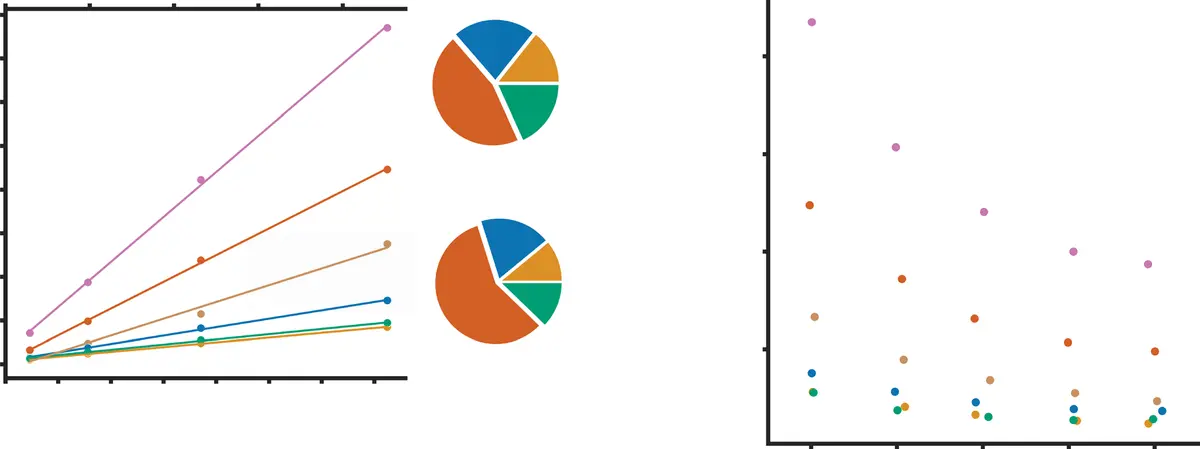
Serial block-face scanning electron microscopy (SBF-SEM) has fundamentally reshaped the landscape of structural biology by permitting the acquisition of three-dimensional tissue ultrastructure at nanoscopic resolution [8, 35, 76, 100, 130, 149, 175]. An automated ultramicrotome mounted within the vacuum chamber of a scanning electron microscope sequentially removes ultrathin sections of a resin-embedded sample, scanning the newly exposed block face iteratively [8, 35, 180]. While this dramatically reduces the z-axis misalignment artifacts common in traditional serial section transmission electron microscopy (ssTEM) [14, 48], the resulting scale of the data is staggering. A single SBF-SEM imaging session can routinely capture volumes spanning hundreds of micrometres [115], yielding complex, highly detailed datasets that range in size from tens of gigabytes to several terabytes—often encompassing tens of thousands of mitochondria or millions of densely packed neurites [130, 149, 175] (Figure 24).

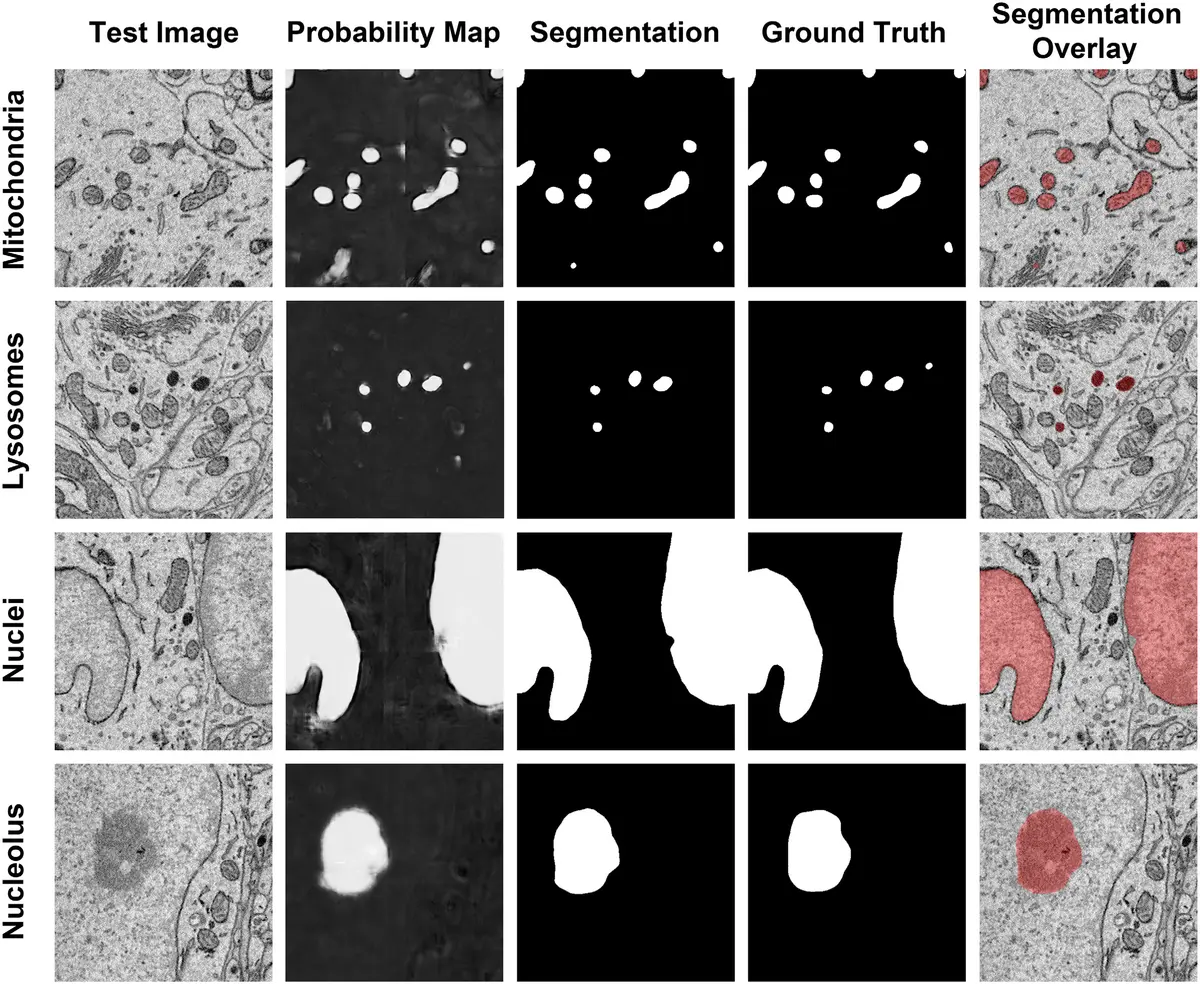
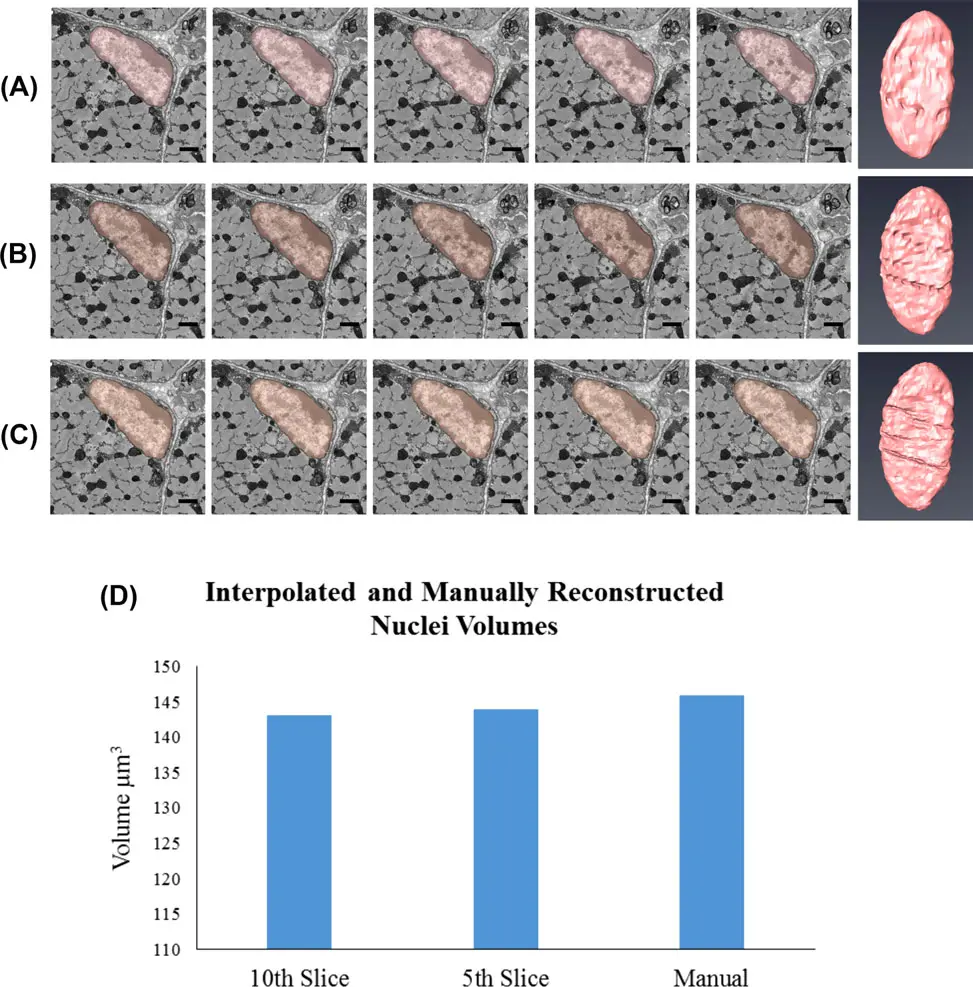
This explosion in data acquisition capability has shifted the primary experimental bottleneck from tissue sectioning and imaging to downstream data processing and analysis [76, 150, 177]. To extract meaningful quantitative morphological metrics or map complex connectomic wiring diagrams, structural elements within the volumetric dataset must be reliably delineated [14, 150]. Historically, this process of image segmentation was achieved through manual annotation, requiring a human expert to trace the boundaries of target structures across thousands of individual two-dimensional slices [14, 100, 150, 171, 175, 177, 180]. Manual segmentation is excruciatingly slow, prone to inter-operator bias, and entirely unscalable; reconstructing a mere 1 mm³ of mammalian cortex would theoretically require over two millennia of continuous human labor [14]. Consequently, the development of computer-aided automated and semi-automated segmentation algorithms has become a paramount priority in the field of volume electron microscopy (vEM) [14, 149, 175] (Figures 25–27).
From Heuristics to Interactive Machine Learning
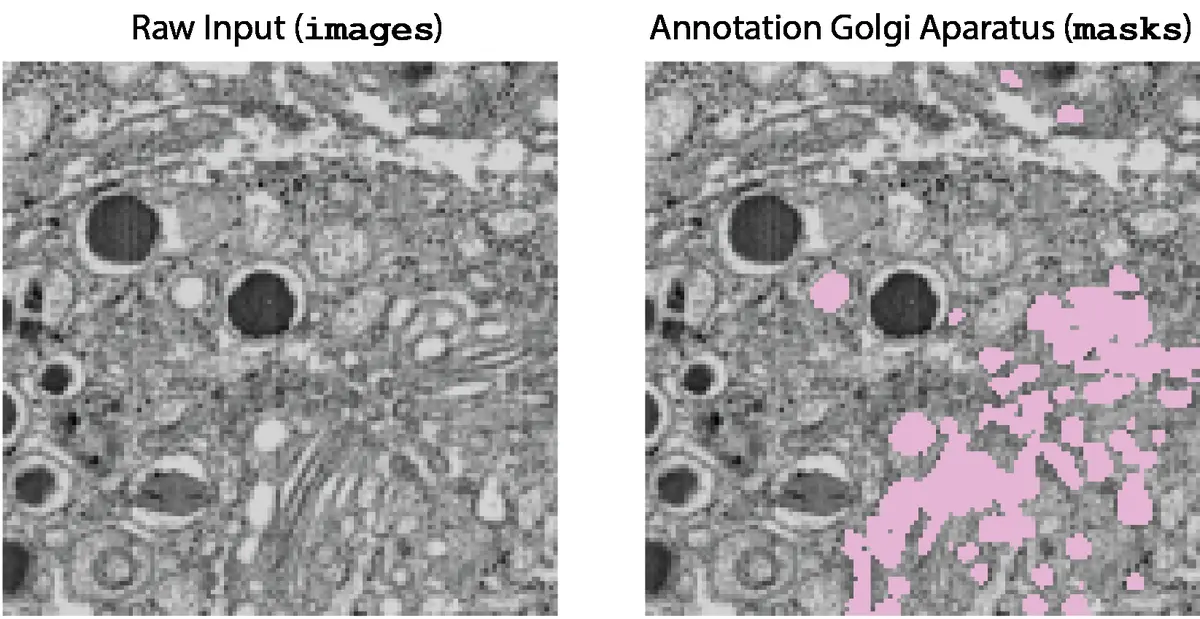
Early attempts to automate the segmentation of vEM datasets relied upon traditional computer vision algorithms such as intensity thresholding, watershed transforms, morphological filtering, and region-growing paradigms [9, 20, 35, 48, 100, 115, 149, 150, 171]. Thresholding assigns voxels to a specific class if their grayscale intensity falls within a predefined range [20, 35, 100, 149]. While highly computationally efficient, these heuristic methods are exceptionally fragile when applied to SBF-SEM data. Electron micrographs lack the distinct multichannel fluorophore targeting of light microscopy; instead, all structures are rendered in a single grayscale channel dependent on the differential uptake of heavy metal stains [8, 76, 170, 172]. Because functionally distinct organelles—such as lysosomes and lipid-rich myelin sheaths—may exhibit nearly identical grayscale intensities, simple thresholding inevitably leads to severe over-segmentation or under-segmentation [76, 172] (Figure 28).
To overcome the limitations of strictly heuristic approaches, the field transitioned toward shallow, interactive machine learning (ML) models [9, 76, 169]. One of the most widespread frameworks in this domain is the Random Forest algorithm [150, 169], popularized by open-source platforms such as *ilastik* [9, 16, 100, 115, 149, 153, 169, 179] and the *Trainable Weka Segmentation* plugin for Fiji/ImageJ [16, 172]. In these interactive workflows, the user provides sparse annotations by "painting" brush strokes over the foreground objects and the background [9, 169]. The software computes a rich array of local image features for every voxel—including Gaussian smoothing for intensity, Hessian eigenvalues for ridges and edges, and structure tensors for texture—which are then fed into the Random Forest classifier [169, 179].
These interactive ML tools offer several advantages: they train rapidly, provide real-time feedback allowing for iterative correction, and demand very little computational power relative to deep learning [169]. However, traditional pixel-classification workflows are inherently limited by their restricted receptive fields. Because they classify voxels based primarily on local texture and edge features without synthesizing broad spatial context, they often fail to capture the complex, long-range morphological characteristics of highly branched organelles like the endoplasmic reticulum, or intertwined neuronal processes [169].
The Deep Learning Revolution: U-Net and Volumetric Architectures
The most profound revolution in SBF-SEM image segmentation has been the advent of artificial intelligence, specifically Deep Convolutional Neural Networks (DCNNs) [16, 162, 169, 170, 171, 174]. Unlike traditional ML models that rely on hand-crafted feature filters, CNNs autonomously learn optimal hierarchical feature representations directly from the training data through backpropagation [162, 169]. Early CNN applications in EM utilized sliding-window patch classification [16, 150], but these were computationally inefficient and lacked global context.
The breakthrough for biomedical imaging came with the introduction of the fully convolutional U-Net architecture [149, 163, 169, 170, 175]. U-Net derives its name from its symmetric, U-shaped design, comprising a contracting (encoder) pathway and an expansive (decoder) pathway [169, 170, 171, 181]. The encoder employs sequential convolutional and pooling layers to compress the spatial dimensions of the image while exponentially increasing the depth of extracted feature channels, effectively capturing deep semantic context [169, 170]. The decoder progressively upsamples this low-resolution feature map to restore the original spatial resolution [170, 171]. Crucially, U-Net utilizes "skip connections" that concatenate the high-resolution feature maps from the encoder directly with the upsampled representations in the decoder [169, 170, 171, 181]. This architectural innovation allows the network to synthesize broad contextual understanding with precise, pixel-level boundary localization, achieving unprecedented segmentation accuracy even with limited training data [169, 170, 175, 181].
While the original 2D U-Net is highly effective, SBF-SEM datasets are inherently volumetric. Processing 3D volumes slice-by-slice discards valuable axial context, which human annotators naturally rely upon to trace continuous structures [149, 175]. To exploit this third spatial dimension, researchers developed 3D variants such as the 3D U-Net [16, 149, 163, 169, 170] and V-Net [149, 163]. These models replace 2D convolutions with 3D convolutional kernels, analyzing contiguous blocks of tissue simultaneously [16, 149, 169]. Although 3D architectures require substantially more graphical processing unit (GPU) memory, they drastically reduce "flickering" or structural discontinuities across the z-axis [149, 175]. Furthermore, highly advanced feature extractors like ResNet and Inception-ResNet-v2 have been integrated into the U-Net encoding pathways [16, 170, 171]. Residual connections in these networks bypass intermediate layers, preventing the vanishing gradient problem and allowing the training of profoundly deep networks capable of abstracting highly complex biological shapes [16, 170, 171].
Recognizing that optimal deep learning architecture depends heavily on the specific dataset, the field has recently gravitated toward self-configuring frameworks like nnU-Net [149, 163, 169]. Rather than requiring manual hyperparameter tuning, nnU-Net automatically analyzes the geometric and intensity properties of the input SBF-SEM dataset and dynamically configures the optimal preprocessing steps, network topology, batch sizes, and loss functions, frequently outperforming manually engineered networks [149, 163, 169].
Instance Segmentation and Connectomics Paradigms
Deep learning in SBF-SEM is broadly categorized into two distinct tasks: semantic segmentation and instance segmentation [149, 171]. Semantic segmentation predicts a class probability for every voxel (e.g., "mitochondrion" versus "background") [149, 171]. However, if two mitochondria are physically touching, semantic segmentation will fuse them into a single continuous blob. Instance segmentation goes a step further by distinctly delineating separate instances of the same class [149, 171].
Instance segmentation is the foundational challenge of "connectomics"—the comprehensive mapping of neuronal circuits [16, 86]. Densely packed axons and dendrites in neural tissue exhibit highly irregular, tortuous paths that span vast distances, rendering standard bounding-box object detection or simple watershed algorithms highly susceptible to split-and-merge errors [16, 171]. To solve this, researchers pioneered Flood-Filling Networks (FFNs) [16, 163, 171, 173, 180]. Unlike standard CNNs that predict all voxels simultaneously in a bottom-up approach, FFNs merge semantic and instance segmentation using a top-down, single-object tracking technique [16, 171]. Employing Recurrent Neural Networks (RNNs), an FFN is seeded at a specific voxel and sequentially "floods" the interior of the cellular process, utilizing previous shape predictions to inform subsequent tracking steps [16, 171]. This maintains the topological continuity of individual neurons over enormous volumes, demonstrating superhuman accuracy on dense circuit reconstructions [16, 163, 171, 180]. Similar multi-object tracking methods utilizing cross-classification clustering have also been successfully deployed [171].
For lower-resolution or large field-of-view SBF-SEM datasets, hybrid geometric pipelines like DeepACSON have been developed [171]. DeepACSON first uses a deep convolutional network to generate semantic probability maps of white matter ultrastructures. To separate touching, under-segmented myelinated axons, it subsequently applies a Cylindrical Shape Decomposition (CSD) algorithm that utilizes the innate tubular geometry of axons as a prior [171]. Similarly, sphericality priors and elastic deformations are used to disentangle adjacent cell nuclei, demonstrating how deep learning can be powerfully combined with traditional geometric constraints [171].
The Ground Truth Bottleneck and Data Augmentation
Despite their unparalleled accuracy, deep learning architectures are notoriously data-hungry, requiring massive amounts of meticulously labeled "ground truth" data for supervised learning [16, 76, 162]. Because generating voxel-perfect annotations in SBF-SEM requires highly specialized domain expertise, acquiring sufficient training data represents the most significant barrier to widespread AI adoption [14, 16, 76, 162, 163, 167, 171].
To circumvent this bottleneck, the bioimage analysis community has embraced multiple strategies. One of the most critical is aggressive data augmentation [16, 149, 171, 181]. By applying stochastic transformations to a small pool of manually annotated images, researchers can artificially inflate the size and diversity of the training set. Common augmentations for SBF-SEM include horizontal and vertical flipping, arbitrary rotations, and shifting [149, 171, 181]. More advanced techniques deploy elastic deformations to simulate the natural biological variance in organelle shape [171], as well as intensity variations, contrast adjustments, and the injection of additive Gaussian noise to mimic varying microscope beam conditions [181]. Some pipelines even intentionally simulate EM-specific imaging artifacts, such as missing z-sections or local misalignments, forcing the network to learn robust, artifact-resistant features [165].
Another revolutionary approach to the annotation deficit is crowdsourcing and citizen science [16, 76, 86, 115, 150]. Platforms like *EyeWire* and *FlyWire* distribute massive unannotated connectomic datasets to thousands of laymen volunteers globally, framing the tracing of neurons as an online puzzle game [16, 86]. The consensus of multiple amateur tracers is aggregated to correct algorithmic errors and generate pristine ground truth [16, 76, 86]. Similarly, distributed collaborative annotation environments such as *CATMAID* and *webKnossos* permit multi-institutional teams of scientists to synchronously skeleton-trace and proofread vast datasets in a shared, cloud-based space [16, 86, 115].
When analyzing inherently sparse structures—such as synaptic clefts or rare vesicular events—the extreme foreground-to-background class imbalance severely destabilizes standard CNN training [16, 167, 171, 181]. If 99% of a volume is background, a network can achieve a 99% accuracy simply by classifying everything as background. To counteract this, dynamic batch sampling is used to heavily favor regions containing the sparse target, and weighted loss functions (such as weighted Dice or cross-entropy loss) are implemented to disproportionately penalize false negatives [181]. Furthermore, some models utilize sparse annotation techniques, predicting dense 3D distance transforms rather than simple binary boundaries, which forces the network to capture broader spatial relationships from very weakly labeled data [16, 167].
Transfer Learning and Domain Adaptation
A persistent challenge in deep learning for SBF-SEM is the problem of "domain shift." An architecture meticulously trained to segment mitochondria in isotropic FIB-SEM mouse brain tissue will frequently fail catastrophically when applied to anisotropic SBF-SEM datasets of rat liver or human HeLa cells [33, 39, 118]. This fragility stems from variations in the physical appearance of organelles across cell types, differences in heavy metal staining protocols, and discrepancies in voxel resolution between different microscopic modalities [162, 163].
To overcome the lack of generalized models, developers increasingly rely on Transfer Learning [162, 163]. Instead of initializing a network with random parameters, transfer learning initializes the network with weights that have already been pre-trained on massive, diverse datasets [162, 163]. Because the initial convolutional layers of a CNN learn fundamental, low-level visual features—such as texture gradients, membranes, and edges—these features are highly conserved across almost all vEM datasets [162, 163]. Once pre-trained, the model undergoes "fine-tuning," where the weights of the deeper, class-specific layers are updated using only a minuscule subset (often just a few slices) of manually annotated data from the target SBF-SEM dataset [162, 163]. This dramatically reduces the time and labor required to segment novel tissue types.
An even more advanced paradigm is Domain Adaptation (DA), which attempts to correct domain shifts without requiring any new ground-truth annotations in the target dataset (unsupervised DA) [33, 39, 104, 118, 135, 153]. DA techniques mathematically align the high-dimensional feature distributions of the source domain (where labels are abundant) with the target domain (where labels are absent) [33, 39, 118]. Several architectures have been proposed for vEM domain adaptation. For example, Y-NET architectures append a secondary, parallel decoder pathway to the standard U-Net [33, 39, 104, 118, 153]. This secondary decoder functions as an auto-encoder, forced to accurately reconstruct both the source and target raw images from the shared latent space [33, 39, 104, 118, 153]. By minimizing the reconstruction loss across both domains simultaneously, the shared encoder is implicitly forced to learn domain-invariant structural representations [33, 39, 118, 153]. Alternatively, Domain-Adversarial Neural Networks (DANN) employ an adversarial classifier that attempts to guess which domain a feature map belongs to; the encoder is optimized to actively fool this classifier, thereby erasing domain-specific artifacts [104, 118, 153].
Generalist Models and the Democratization of AI Tools
The culmination of transfer learning and massive data aggregation has led to the recent emergence of highly robust "generalist" models, echoing the "Segment Anything" paradigms seen in natural image processing [149, 163]. Instead of relying on bespoke, single-use algorithms, laboratories can now leverage out-of-the-box foundation models.
A prominent example is *CellPose*, a generalist deep learning architecture initially developed for fluorescence microscopy but highly adaptable to SBF-SEM via transfer learning [9, 124, 162, 167]. CellPose replaces the traditional binary boundary predictions with vector gradient fields [124, 162, 167]. The network predicts topological maps that simulate a physical "flow" pointing toward the geometric center of every individual object [162]. This vector flow inherently resolves touching objects and achieves outstanding instance segmentation. Researchers have demonstrated that fine-tuning a pre-trained CellPose model on a fraction of SBF-SEM platelet and thrombi images yields near-perfect segmentation of densely packed, complex cellular domains where base models previously failed entirely [162].
Similarly, *MitoNet* (often accessed via the Empanada plugin in Napari) provides a generalized convolutional architecture explicitly trained on a massive and heterogeneous corpus of electron microscopy imagery containing over 1.5 million bounding boxes and masks [163, 178]. MitoNet is capable of highly accurate mitochondrial segmentation across radically different tissue types—from HeLa cells to rat pancreas—frequently in a "zero-shot" capacity, meaning it requires zero fine-tuning on the user's local dataset [163, 178]. The *Segment Anything Model 2 (SAM2)* has also been introduced to vEM; pre-trained on billions of masks across diverse visual media, SAM2 utilizes promptable bounding boxes or sparse click coordinates to instantly generate volumetric segmentation masks across an SBF-SEM z-stack [149, 163].
For these highly advanced models to yield a tangible impact, they must be accessible to biologists who lack computational or coding expertise [9, 19, 35]. To bridge this usability gap, the community has developed a rich ecosystem of graphical user interfaces and cloud-based platforms [9, 19, 35]. *DeepMIB* (Microscopy Image Browser) provides a comprehensive, open-source MATLAB environment integrating data alignment, manual curation, and deep learning network training in a single workflow [9, 16, 28, 81, 115, 171]. To mitigate the immense hardware cost of GPU arrays, cloud-computing paradigms like *CDeep3M* have been introduced, allowing researchers to upload SBF-SEM data to remote clusters where pre-configured state-of-the-art CNNs execute the segmentation natively in the cloud [9, 16, 149, 176]. Furthermore, tools such as *ZeroCostDL4Mic* [9, 16, 149] and *DeepImageJ* [16, 149] integrate seamlessly into familiar platforms like ImageJ/Fiji. These tools provide pre-written Jupyter notebooks running on free Google Colab servers, fully democratizing access to supercomputing infrastructure.
By integrating U-Net architectures, self-configuring pipelines like nnU-Net, generalist transfer-learning models, and user-friendly open-source software, the analysis of SBF-SEM datasets is transitioning from a prohibitive, multi-year manual endeavor to a highly streamlined, automated computational pipeline.