Comparison with FIB-SEM and Other Volume EM Techniques
The rapid evolution of volume electron microscopy (vEM) has fundamentally transformed our ability to investigate the three-dimensional (3D) ultrastructure of cells, tissues, and whole organisms [9, 14, 27, 143, 157]. Within the diverse landscape of vEM, modalities are broadly delineated into two categories: serial sectioning approaches, which collect physical tissue slices prior to imaging, and block-face approaches, which repeatedly image the surface of a sample as it is sequentially ablated [9, 18, 22, 80]. As a premier block-face technique, serial block-face scanning electron microscopy (SBF-SEM) occupies a critical functional niche. By integrating an ultramicrotome directly into the vacuum chamber of a scanning electron microscope (SEM), SBF-SEM automates the slice-and-view cycle, offering an optimal balance of throughput, field of view (FOV), and resolution [5, 13, 18, 73]. However, no single vEM technique represents a universal solution. To fully appreciate the capabilities and limitations of SBF-SEM, it must be critically compared against other leading vEM technologies, most notably focused ion beam scanning electron microscopy (FIB-SEM), serial section transmission electron microscopy (ssTEM), and automated tape-collecting ultramicrotome SEM (ATUM-SEM) (Figures 63–66).
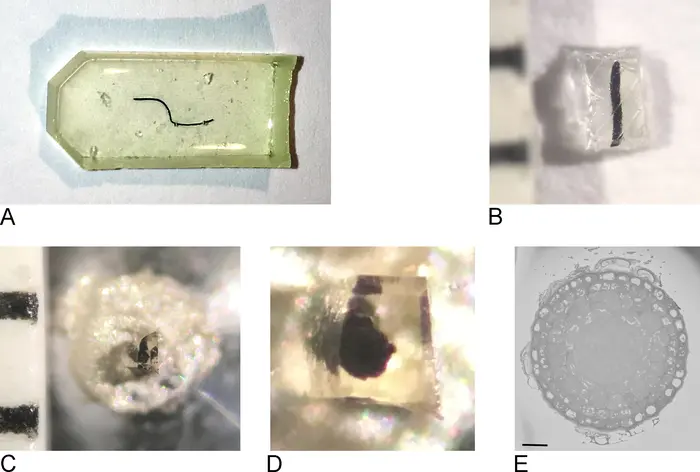

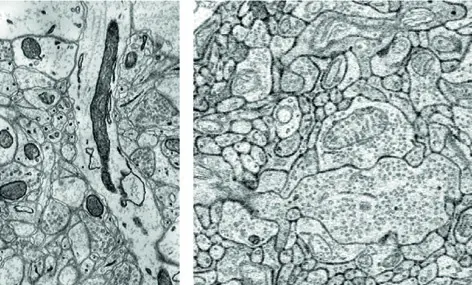
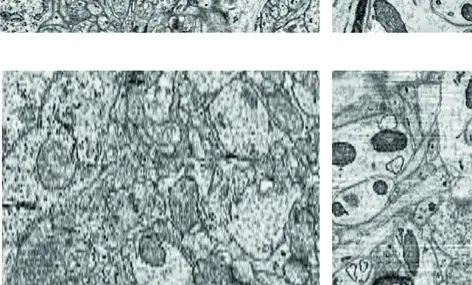
SBF-SEM versus Focused Ion Beam Scanning Electron Microscopy (FIB-SEM)
SBF-SEM and FIB-SEM share a fundamental operational paradigm: both are destructive, block-face scanning techniques that acquire serial images using backscattered electron (BSE) detectors to generate heavy-metal-dependent contrast [5, 9, 27, 38, 42, 73]. Their primary divergence lies in the mechanism of tissue ablation, which inherently dictates their respective spatial resolutions and practical imaging volumes.
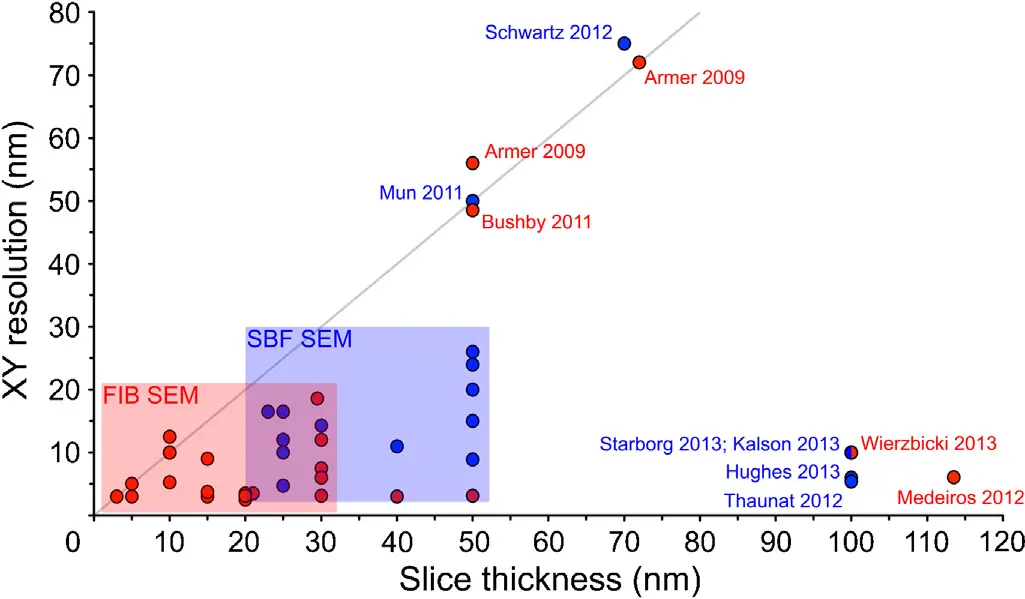
**Resolution and Isotropic Imaging** In SBF-SEM, the sample surface is removed mechanically using a diamond knife [9, 37, 38, 73]. The axial (z) resolution is therefore physically constrained by the minimum thickness of the resin slice that can be reliably cut without causing intolerable sectioning artefacts, tearing, or severe compression [26, 32, 111]. For most biological specimens, this functional limit is approximately 25 to 50 nm [18, 26, 32, 42, 50]. Because the lateral (x, y) resolution of the SEM can easily reach 3 to 5 nm, SBF-SEM datasets are characteristically anisotropic [14, 18, 38, 266, 284]. This anisotropy can complicate the precise 3D rendering and semi-automated segmentation of minute or highly complex subcellular architectures, such as mitochondrial cristae, small vesicles, or fine dendritic spines [14, 28, 154, 266].
FIB-SEM, conversely, employs a focused beam of gallium ions (or increasingly, plasma ions like xenon or oxygen) to mill away the sample surface [9, 16, 38, 73]. Because ion milling is not bound by the mechanical limitations of a physical blade, it can ablate arbitrarily thin layers of material [9, 20, 32, 42]. This permits FIB-SEM to achieve exceptional, isotropic 3D resolution, routinely generating voxel dimensions of 3 to 5 nm in all axes [16, 18, 26, 38, 72, 140, 143, 148, 284]. Consequently, FIB-SEM is universally considered the superior technique for the detailed evaluation of fine ultrastructure at the single-cell or organelle level [16, 38, 266, 284].
**Field of View and Throughput** The unparalleled z-resolution of FIB-SEM comes at the significant expense of field of view and throughput. Ion beam milling is an inherently slow, highly localised process. To prevent milling aberrations such as "curtaining," the region of interest must remain relatively small, restricting the practical FOV to tens of micrometres (e.g., 20 to 50 µm) [14, 16, 18, 32, 38, 42, 72]. Consequently, the total volume effectively interrogated by conventional FIB-SEM typically ranges from to [14, 16, 26, 32, 42].
SBF-SEM excels precisely where FIB-SEM is limited. The mechanical stroke of a diamond knife can clear a massive surface area in mere seconds, allowing SBF-SEM to effortlessly sweep across broad fields of view (up to ~800 or more) [5, 14, 18, 20, 26, 32, 38, 266]. This unmatched slicing speed permits the rapid acquisition of macroscopic tissue volumes exceeding to , making SBF-SEM the workhorse for tissue-level connectomics, vascular mapping, and developmental biology studies where establishing the spatial context between numerous cells is paramount [14, 16, 20, 26, 32, 42, 72].
**Sample Integrity and Hybrid Workflows** The interaction between the electron beam and the non-conductive embedding resin presents persistent challenges for block-face techniques. SBF-SEM is particularly vulnerable to charge accumulation in areas of bare resin, which can induce severe image warping, beam drift, and knife chatter [8, 29, 32, 38, 44, 77, 111]. Addressing this requires intense *en bloc* heavy metal staining protocols and often the use of variable-pressure chamber environments or focal charge compensation (nitrogen gas injection), which can inadvertently degrade lateral resolution [29, 32, 77, 140]. Interestingly, FIB-SEM is somewhat less susceptible to these widespread charging artefacts because the continuous implantation of positive gallium ions (Ga+) during milling acts to neutralise the negative charge deposited by the electron beam [5, 38].
Because SBF-SEM physically destroys the entire cross-section of the block during each cut, the exact region of interest must be selected blindly from the first z-stack image [22, 29, 32, 87]. If a specific, rare cellular event is missed, the entire sample is lost. FIB-SEM, however, only destroys the local trench being milled, preserving the surrounding tissue block for subsequent sequential acquisitions [28, 32, 92]. To exploit the strengths of both modalities, researchers have developed highly efficient synergistic workflows: SBF-SEM is first utilised at low magnification to rapidly screen vast tissue volumes to locate a rare structure (e.g., a specific pericyte or synaptic convergence), after which the sample is transferred to a FIB-SEM to mill and image the targeted event at supreme isotropic resolution [5, 28]. Furthermore, innovations such as multi-energy deconvolution—where sequentially varied accelerating voltages computationally extract depth information from the block-face—are actively bridging the z-resolution gap, allowing SBF-SEM to push toward isotropic 10 nm voxels [14, 18, 42, 143].
SBF-SEM versus Serial Section TEM (ssTEM) and Electron Tomography (ET)
For decades prior to the advent of automated block-face SEM, three-dimensional ultrastructural analysis relied exclusively on serial section transmission electron microscopy (ssTEM) and serial section electron tomography (ssET) [14, 18, 37, 73]. SBF-SEM emerged as a direct technological response to the severe logistical limitations of these traditional TEM methodologies [13, 35, 73].
**Resolution Profiles** In ssTEM, ultrathin sections (typically 50–70 nm) are physically cut, floated onto a water bath, and manually collected onto film-coated grids for imaging via transmitted electrons [9, 14, 72, 73, 284]. Because TEM operates at high accelerating voltages (e.g., 80–300 kV) and relies on transmitted rather than backscattered electrons, ssTEM consistently achieves the highest absolute lateral spatial resolution, easily reaching sub-nanometre scales [9, 14, 16, 38, 284]. In contrast, the lateral resolution of SBF-SEM is generally restricted to 3–10 nm due to the larger interaction volume of the primary electron beam and the reliance on BSE signals [14, 16, 38, 50, 67, 284].
To overcome the poor axial resolution inherent to standard 2D sections, serial section electron tomography (ssET) tilts relatively thick sections (200–1000 nm) through a high angular range (e.g., ±70 degrees) inside the TEM, computationally back-projecting the images to yield exquisite 3D volumes with ~2 nm z-resolution [9, 14, 17, 18, 20]. However, ssET is severely handicapped by the "missing wedge" of data lost at extreme tilt angles, a highly restricted FOV (often just a few micrometres), and painstaking acquisition times [14, 17, 18, 20, 143].
**Automation, Labour, and Geometric Distortion** The most critical advantage of SBF-SEM over ssTEM is the radical reduction in manual labour and the elimination of section handling artefacts. ssTEM requires the meticulous, highly skilled manual collection of hundreds of fragile serial sections [13, 14, 20, 87, 92, 141]. The technique is notoriously prone to catastrophic errors: a single lost section, a fold, stretching, physical compression, or heavy metal stain precipitation can permanently sever the continuity of a 3D reconstruction [13, 14, 20, 35, 73, 87, 92, 111]. Furthermore, each physical section deforms slightly upon water contact and grid mounting. Consequently, assembling an ssTEM volume requires mathematically complex, computationally heavy non-linear alignment algorithms to correct image distortions and register the stack [13, 14, 35, 73, 92].
SBF-SEM entirely bypasses these mechanical pitfalls. Because the backscattered electron image is captured directly from the rigid, bulk resin block-face *prior* to the physical removal of the section, section loss is irrelevant, and the physical deformation of the slice has no impact on the acquired data [13, 18, 72, 73, 111]. As a result, SBF-SEM datasets are inherently pre-aligned. They exhibit minimal geometric distortion and typically require only basic, rigid translational registration (e.g., cross-correlation or simple SIFT algorithms) to correct for minor beam drift or charging deflections [14, 18, 26, 28, 73, 87, 111]. Once the sample is properly mounted and imaging conditions are established, SBF-SEM instrumentation can operate autonomously for days or weeks, massively reducing the human capital required to map large volumes [13, 18, 20, 37, 73, 111].
SBF-SEM versus Array Tomography (AT) and ATUM-SEM
To merge the high-throughput capabilities of SEM with the non-destructive advantages of serial sectioning, researchers developed Array Tomography (AT) and its highly automated successor, the Automated Tape-collecting Ultramicrotome (ATUM-SEM) [9, 16, 27, 36, 141]. In AT, long ribbons of serial ultrathin sections are collected onto solid substrates (such as silicon wafers, glass slides, or ITO-coated coverslips) [2, 9, 36]. ATUM scales this process by capturing thousands of consecutive sections directly from the diamond knife onto a continuously spooling conductive Kapton tape, which is subsequently segmented and mounted onto wafers for SEM imaging [9, 16, 72, 73, 141].
**The Paradigm of Sample Preservation** The fundamental dichotomy between SBF-SEM and ATUM-SEM is the fate of the physical sample. SBF-SEM is intrinsically and irreversibly destructive [18, 22, 29, 73, 87, 111]. The tissue is permanently obliterated by the microtome during acquisition; sections cannot be archived, and the block cannot be re-examined to clarify ambiguities or image a missed structure [22, 29, 73, 87, 111].
Conversely, ATUM-SEM and Array Tomography generate a permanent, physical tissue library [16, 141]. Because the ultrathin sections are safely preserved on the substrate, researchers can employ powerful hierarchical, multi-scale imaging strategies [16, 73, 141]. An investigator can rapidly scan the entire tape at low magnification to construct a mesoscopic roadmap of the tissue, identify specific anatomical ROIs, and then selectively direct the SEM to re-image only those precise targets at nanometre resolution [16, 73, 141].
**Correlative Light and Electron Microscopy (CLEM)** The preservation of sections in AT also provides unparalleled advantages for Correlative Light and Electron Microscopy (CLEM). Because the sections rest on a stable substrate, they can be subjected to post-embedding immunofluorescence and imaged with an optical or super-resolution fluorescence microscope prior to heavy-metal contrasting and SEM imaging [2, 3, 36, 44, 154]. This allows researchers to directly map molecular identities (via fluorescent antibodies) to precise ultrastructural coordinates on the exact same tissue slice [2, 36, 44, 154]. SBF-SEM is highly hostile to standard CLEM workflows; because the tissue is imaged in bulk under vacuum and subsequently destroyed, correlation usually requires complex pre-embedding fluorescent markers (e.g., APEX2), reliance on endogenous fiducials (like blood vessels), or the laborious use of near-infrared (NIR) laser branding to burn reference marks into the tissue block to guide the SBF-SEM to the ROI [3, 50, 77, 140, 185].
**Parallelisation and Acquisition Throughput** While SBF-SEM boasts excellent automation, its throughput is strictly limited by the sequential bottleneck of the machine: a single electron beam must slowly raster across the block-face before the knife can act [5, 10, 20, 266]. ATUM-SEM decouples the sectioning process from the imaging process. Once the sections are collected on tape, they can be distributed across multiple standard SEMs operating simultaneously, or more profoundly, they can be imaged using revolutionary multibeam SEM (mSEM) technology [9, 10, 16, 22, 72, 178]. By utilising 61 or more electron beamlets to image a section concurrently, mSEM combined with ATUM facilitates data acquisition rates that eclipse SBF-SEM by orders of magnitude, making it the preferred pipeline for massive, petabyte-scale connectomics projects (such as mapping entire fly or mouse brains) [10, 16, 178].
**Data Processing and Alignment** Despite its immense throughput potential, ATUM-SEM suffers from the same fundamental data processing handicap as ssTEM. The physical collection of sections onto tape inevitably induces stretching, wrinkling, and non-linear deformations [9, 14, 73]. To reconstruct a coherent 3D volume, ATUM datasets demand sophisticated, computationally punishing alignment algorithms—such as elastic volume reconstruction or SIFT-based global minimisation frameworks—to digitally unwarp the tissue [9, 14, 73, 141]. SBF-SEM, by avoiding section collection entirely, remains vastly superior for laboratories seeking to minimise the computational overhead of alignment, delivering robust, natively registered volumes straight from the microscope [13, 72, 73, 87, 111].
Summary and the Future of Volume EM
The selection of a vEM modality represents a careful compromise between spatial resolution, field of view, acquisition speed, sample preservation, and computational infrastructure [16, 38, 157, 266, 284]. FIB-SEM delivers unmatched isotropic resolution ideal for mapping the internal architecture of single cells and small organelles, but is severely limited in its volumetric reach [16, 38, 42, 284]. ssTEM provides the absolute highest lateral resolution but is bottlenecked by manual labour and extreme alignment challenges [14, 20, 92]. ATUM-SEM and Array Tomography offer non-destructive tissue libraries, direct CLEM capabilities, and massive parallelisation potential, but require significant computational power to correct sectioning distortions [14, 16, 36, 141].
Within this matrix, SBF-SEM occupies a highly strategic and powerful niche. It provides a robust, fully automated "slice-and-view" pipeline that bridges the gap between the mesoscopic tissue context and nanoscopic cellular detail [18, 26, 28, 37, 72, 111]. While its anisotropic z-resolution (~25 nm) and destructive nature preclude the finest isotropic rendering or post-acquisition molecular probing, SBF-SEM's ability to rapidly and reliably generate massive, pre-aligned 3D ultrastructural datasets makes it an indispensable tool [12, 28, 87, 111]. As the field progresses, the integration of SBF-SEM into multiscale, multi-modal workflows—such as combining it with X-ray micro-CT for initial targeting or FIB-SEM for high-resolution deep-dives—will continue to expand our understanding of complex biological systems [5, 9, 38]. The ultimate bottleneck for SBF-SEM, as with all vEM techniques, is rapidly shifting from data acquisition to the formidable challenge of AI-driven image segmentation and data management [9, 14, 16, 163].