Disease and Pathology
Introduction to 3D Ultrastructural Pathology
Historically, the structural analysis of diseased tissues has relied heavily on transmission electron microscopy (TEM) to uncover subcellular pathology at the nanometre scale. However, the two-dimensional (2D) nature of conventional TEM provides only a restricted snapshot of inherently three-dimensional (3D) biological environments, which can obscure complex topological changes induced by disease [9, 10, 34]. Serial block-face scanning electron microscopy (SBF-SEM) has emerged as a transformative technique that bridges the significant resolution and volume gap between optical fluorescence microscopy and high-resolution electron tomography [17, 33, 37, 100, 268]. By iteratively coupling the removal of ultrathin resin sections with backscattered electron imaging of the block face, SBF-SEM routinely generates highly aligned, large-volume 3D datasets ( to ) without the manual distortions and limitations associated with serial section TEM [11, 17, 32, 33, 34, 72]. This automated capability permits the exhaustive characterisation of entire cells, tissue microenvironments, and rare pathological anomalies, fundamentally reshaping our understanding of disease mechanisms across diverse organ systems [10, 22, 29, 55].
Renal Pathology and Glomerular Disease
The application of SBF-SEM in nephrology has yielded unprecedented insights into the 3D architecture of the kidney, particularly the intricate filtration barrier of the glomerulus. In healthy states, SBF-SEM allows for the precise volumetric reconstruction of podocytes, mapping the complex arborisation of their major processes and interdigitating foot processes [40, 68]. Under pathological conditions, this technique has revealed previously unappreciated ultrastructural aberrations. For instance, SBF-SEM imaging of autoimmune nephropathy models has highlighted regional immune deposits, heterogeneous thinning, and double contouring of the glomerular basement membrane (GBM) [107]. Furthermore, 3D ultrastructural imaging has been utilised to examine glycosaminoglycan deposition within kidney glomeruli, providing new insights into filtration barrier integrity [12].
Perhaps the most striking discoveries facilitated by SBF-SEM in renal pathology relate to the physical interactions between resident glomerular cells during severe inflammation. In comprehensive 3D analyses of human biopsy specimens from patients with lupus nephritis and IgA nephropathy, SBF-SEM uncovered massive structural reorganisations that were virtually invisible in 2D sections [9, 264, 265]. Researchers successfully tracked podocyte cytoplasmic processes as they profoundly disrupted and invaginated into the GBM [9, 265]. In regions of mesangial interposition, podocytes penetrated the subendothelial area, extending highly complex, spike-like protrusions through the GBM to make direct cytoplasmic contact with invading mesangial cells [264]. In a single reconstructed lupus nephritis tissue volume, a penetrating podocyte was shown to form over 100 distinct contact sites with surrounding mesangial cells [9, 264]. These unique structural communications represent a common, yet previously hidden, feature of podocyte behaviour in response to severe glomerular inflammation [264]. Similar GBM abnormalities and active podocyte protrusions—resembling invadosomes—have also been identified in genetic murine models of Alport syndrome and focal segmental glomerulosclerosis, suggesting that podocyte-matrix adhesion pathways are universally activated following primary molecular insults to the filtration barrier [270].
Liver Pathology and Hepatocellular Alterations
In hepatology, SBF-SEM has been rigorously applied to unravel the 3D microarchitecture of liver tissue, enabling the simultaneous quantification of parenchymal cells, Kupffer cells, the anastomosing network of hepatic sinusoids, and bile canaliculi [34]. Comprehensive appraisals of varied sample preparation protocols have confirmed that combining heavy metal fixatives, stains, and mordanting agents provides the necessary contrast to accurately reconstruct hepatocellular ultrastructure, including mitochondria, rough endoplasmic reticulum, glycogen, and lipid droplets, without charging artefacts [34]. Correlative light and electron microscopy (CLEM) workflows have further augmented this approach, enabling the targeted 3D tracking of specific subcellular events, such as lipid droplet accumulation and cytoskeletal changes, which are central to the pathogenesis of fatty liver disease and toxic injury [34].
SBF-SEM has also solved longstanding morphological challenges regarding specific organelles, such as the Golgi apparatus. Historically defined by 2D TEM criteria, the true holistic geometry of the hepatic Golgi network is difficult to assess [154]. By combining SBF-SEM with zinc-iodine-osmium (ZIO) histochemistry, researchers have successfully reconstructed the Golgi apparatus within whole rat hepatocytes without the need for additional conductive staining [154]. This approach revealed that plural Golgi networks exist within a single hepatocyte, featuring variable cis/trans axis orientations and a unique topographic predilection for regions near the nucleus and bile canaliculi [154]. As artificial intelligence and deep-learning software streamline the classification of these structures, tracking whole-cell Golgi morphology will become a vital metric for elucidating the pathogenesis of hepatic metabolic and secretory diseases [154].
Cancer Cell Ultrastructure and the Tumor Microenvironment
The oncology field has rapidly integrated volume electron microscopy to map the dense, heterogeneous architecture of the tumour microenvironment. Understanding the spatial relationship between cancer cells, stromal components, and vascular networks is essential for deciphering the mechanisms of tumour progression, metastasis, and therapeutic resistance [10, 39, 134]. SBF-SEM analysis of patient-derived xenograft (PDX) models of hepatoblastoma, a common paediatric liver cancer, has revealed unique 3D capillary arrangements and bile canaliculus-like structures, leading to the hypothesis that local capillary architecture directly dictates the ultrastructural orientation of surrounding tumour cells [9, 10, 134]. By coupling SBF-SEM with semi-automated machine learning, researchers have extracted precise bioarchitectural parameters, quantifying the 3D distance between cancer cell nuclei and the nearest blood vessels [9, 134].
In metastatic breast cancer, multi-omic analyses paired with volume EM have highlighted distinct survival mechanisms. 3D reconstructions have visualised tumour cells extending highly organised projections into the host stroma, alongside orderly mitochondrial arrangements, active lamellipodia, macropinosomes, and densely packed clusters of lysosomes, exposing novel structural vulnerabilities [10, 39]. Similar ultrastructural characterisation has been performed on pancreatic ductal adenocarcinoma (PDAC) metastases in the liver, detailing the complex interface between the tumour and the host microenvironment [266]. Additionally, in the diagnosis of papillary thyroid carcinoma, SBF-SEM has proven invaluable for accurately differentiating between "open" intranuclear cytoplasmic inclusions (which maintain clear topological connections to the extranuclear cytoplasm) and "closed" inclusions, providing mechanistic insights into their formation [10].
SBF-SEM has also been instrumental in validating *in vitro* oncological models. A persistent challenge in cancer research is that 2D monolayer cultures often fail to replicate the true physiology of *in vivo* solid tumours. Comparative SBF-SEM analyses of cancer cells grown as traditional monolayers versus 3D "tumorspheres" have uncovered stark ultrastructural disparities [61]. Cells within tumorspheres exhibit a highly complex morphology, including an increased prevalence of deep nuclear envelope invaginations, highly diverse mitochondrial networks, and intricate cellular interconnectivity that is completely absent in 2D cultures [61]. These findings structurally validate tumorspheres as a vastly superior and more physiologically relevant model for cancer drug screening and personalised medicine [61].
Neuropathology and Neurodegenerative Disease
The neurosciences were among the earliest adopters of SBF-SEM, driven by the need to map extensive axonal and dendritic arbours with sufficient resolution to identify individual synaptic vesicles [14]. Beyond basic connectomics, SBF-SEM is now pivotal in characterising neuropathology. Retinal degenerative conditions, for example, often manifest subtle, pre-clinical structural abnormalities before the onset of overt blindness. By automating the capture of massive retinal volumes, SBF-SEM has successfully mapped the 3D architecture of rod and cone photoreceptors, internal disc elements, and the retinal pigment epithelium, revealing isolated, precipitating phenotypic events in mouse models of human retinal dystrophies [55, 95]. In models of diabetic retinopathy, SBF-SEM has elucidated previously uncharacterised 3D changes within the retinal neurovascular unit, including the quantification of pericyte-endothelial peg-and-socket junctions, endothelial tubule formation, and vascular basement membrane thickening [267].
Neurodegenerative and psychiatric disorders are frequently accompanied by widespread subcellular failure. In models of schizophrenia and intellectual disability (such as Schnurri-2 knockout mice), SBF-SEM has been utilised to identify immature morphological properties and subcellular-scale structural defects in the dentate gyrus [85]. Following traumatic brain injury, automated 3D morphometry has demonstrated that myelinated axons in the corpus callosum suffer significant chronic reductions in diameter and exhibit highly irregular, elliptic geometries rather than uniform circular profiles [115]. Furthermore, because SBF-SEM excels at resolving the complex geometry of mitochondria [111, 199], it has become a premier tool for mapping mitochondrial network fragmentation, enlargement, and spatial maldistribution in models of Parkinson’s, Alzheimer's, Huntington's, and amyotrophic lateral sclerosis (ALS) [111, 199].
The technique has also profoundly advanced our understanding of demyelinating diseases such as multiple sclerosis (MS). SBF-SEM permits the precise differentiation of central nervous system (CNS) cells based on ultrastructural criteria—such as nuclear shape, mitochondrial distribution, and the presence of osmiophilic granules [199, 205, 209]. In experimental autoimmune encephalomyelitis (EAE) mouse models, SBF-SEM proved that early demyelination is driven almost entirely by infiltrating monocyte-derived macrophages, while resident microglia remain largely uninvolved during the initial inflammatory stages [199, 209]. In models of chronic demyelination, volumetric imaging has further revealed a decreased number, yet paradoxically increased volume, of cerebellar synaptic terminals, coupled with significant mitochondrial enlargement [8].
Host-Pathogen Interactions and Infectious Disease
Volume electron microscopy is uniquely suited to capturing the spatial dynamics of infectious diseases. SBF-SEM has been adapted to study the life cycles of various pathogens within tissue culture monolayers, circumventing the limitations of 2D sectioning to capture entire viral assembly and egress pathways [26]. For example, 3D reconstructions of sympathetic ganglia infected with alpha-herpesviruses have revealed the striking formation of nuclear actin filaments—a structural manipulation of the host cell entirely driven by the viral infection [269].
SBF-SEM has also been instrumental in defining the profound neurological consequences of parasitic infections. *Toxoplasma gondii*, an intracellular parasite that establishes chronic infection in the mammalian brain, is known to cause severe neuronal changes. Using SBF-SEM to reconstruct layers of the cerebral cortex and hippocampus in infected mice, researchers documented a dramatic immune response characterised by the physical ensheathment of neurons by myeloid-derived cells (primarily reactive microglia and infiltrating macrophages) [203, 205]. In some instances, a single infected neuron was observed to be completely encapsulated by three distinct phagocytic cells, which extended complex, interlocking processes around the neuronal soma [203, 205] (Figure 56).
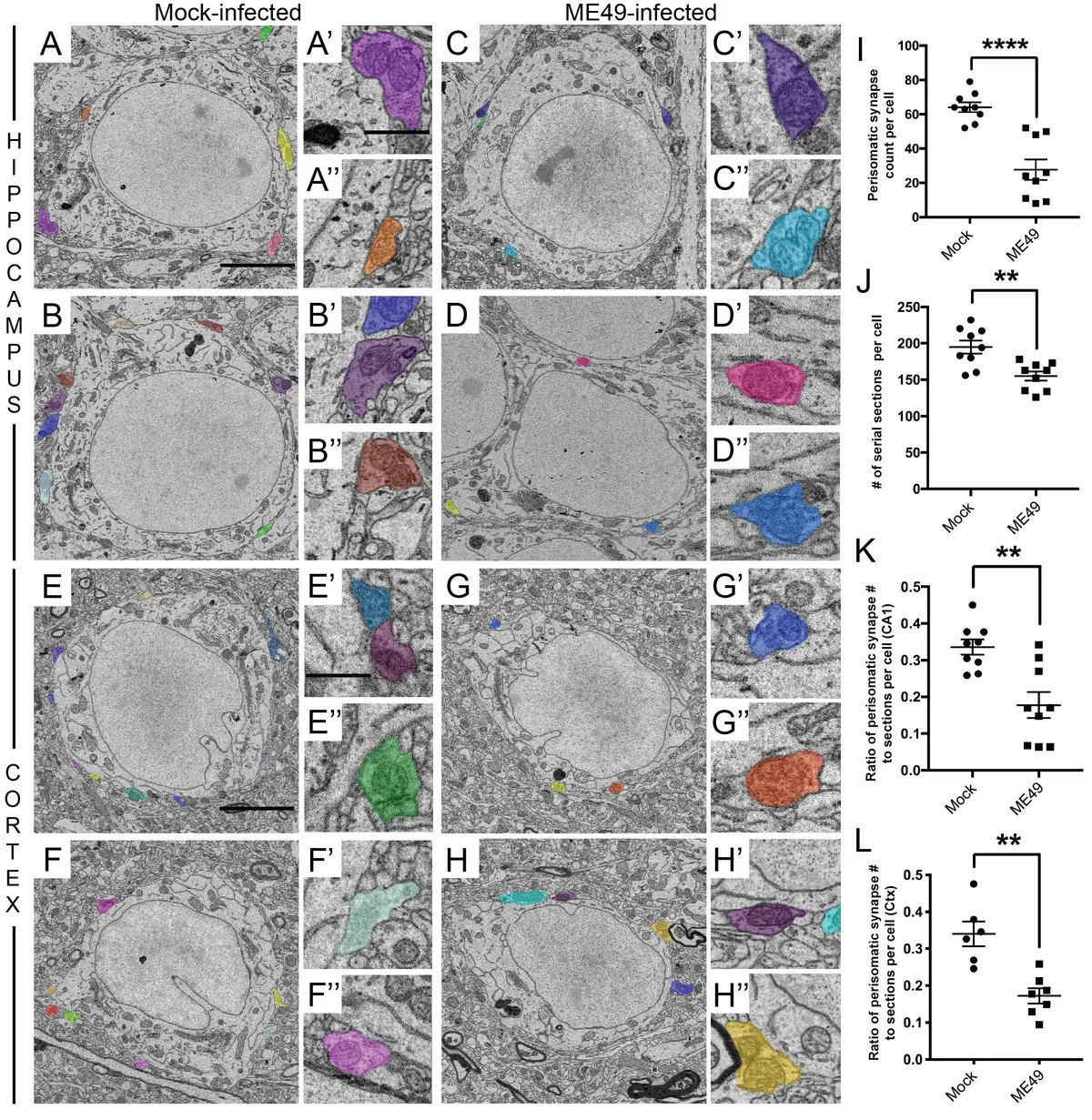
Crucially, SBF-SEM demonstrated that this microglial ensheathment actively drives the stripping of perisomatic inhibitory synapses [203, 205]. By exhaustively tracing pre-synaptic terminals (characterised by synaptic vesicles and active zones opposing the soma), researchers quantified a highly significant loss of GABAergic (GAD67-positive) nerve terminals surrounding the neurons in *Toxoplasma*-infected brains [203, 205]. This large-scale synaptic stripping provides a direct structural explanation for the altered neurocircuitry, hyperexcitability, and behavioural changes frequently observed in chronic toxoplasmosis [203, 205].
Diagnostic Potential and Clinical Translation
While the integration of volume EM into clinical practice presents certain ethical, logistical, and computational hurdles [10], its diagnostic potential is immense. Traditional diagnostic EM relies heavily on 2D TEM for the evaluation of renal biopsies and ciliopathies [9]. However, vEM techniques have already demonstrated superiority in detecting features that escape 2D analysis. For example, in primary ciliary dyskinesia, 3D imaging can detect volumetric reductions in proximal outer dynein arms that are invisible on standard ultrathin sections [9]. In renal pathology, as previously noted, SBF-SEM is required to fully appreciate the extent of GBM disruption and the complexity of cell-cell contacts in lupus nephritis [9, 264].
A primary barrier to the clinical adoption of SBF-SEM has been the extensively long, highly specialised, and toxic sample preparation protocols historically required for whole-block conductivity and contrast [11]. To address this, rapid and simplified specimen preparation methods have been developed specifically for clinical throughput [107]. By employing a modified reduced osmium–thiocarbohydrazide–osmium (rOTO) staining protocol combined with conductive resins, researchers can now bypass the need for radioactive uranyl acetate while still achieving excellent membrane contrast and preventing electron charging artefacts [45, 107, 264]. This streamlined workflow enables the preparation of human tissue biopsies—such as those from chronic kidney disease patients—in a single day, drastically reducing costs and turnaround times [107, 264].
As the speed of automated image acquisition accelerates and data-storage capabilities expand, the final remaining bottleneck in clinical SBF-SEM is image segmentation [16, 35, 100, 264]. The manual tracing of cellular contours across thousands of serial images is prohibitive for routine diagnostics. However, the rapid development of deep-learning algorithms, transfer learning, and AI-based image recognition promises to fully automate the classification of organelles and pathological lesions [10, 134, 154, 163, 264]. By combining rapid, uranyl-free preparation protocols with AI-driven segmentation, SBF-SEM is poised to transition from a highly specialised research tool to a routine, powerful modality in diagnostic surgical pathology [107, 264].