Developmental Biology and Embryology
Bridging the Resolution Gap in Developmental Biology
Understanding the intricate processes of embryonic development requires imaging modalities capable of resolving cellular and subcellular events within their native three-dimensional (3D) tissue environment [12, 13]. Historically, developmental biologists faced a difficult compromise between light microscopy and standard transmission electron microscopy (TEM) [12]. Multiphoton and confocal fluorescence microscopy permit live imaging deep into tissues, but lack the spatial resolution necessary to resolve fine ultrastructural details such as transport vesicles, individual organelles, and extracellular matrix (ECM) assemblies [12]. Conversely, serial section TEM (ssTEM) provides unparalleled nanometer-scale resolution but is highly labor-intensive, restricted to relatively small tissue volumes, and plagued by the physical distortion or loss of fragile ultrathin sections during manual collection [12, 13, 35].
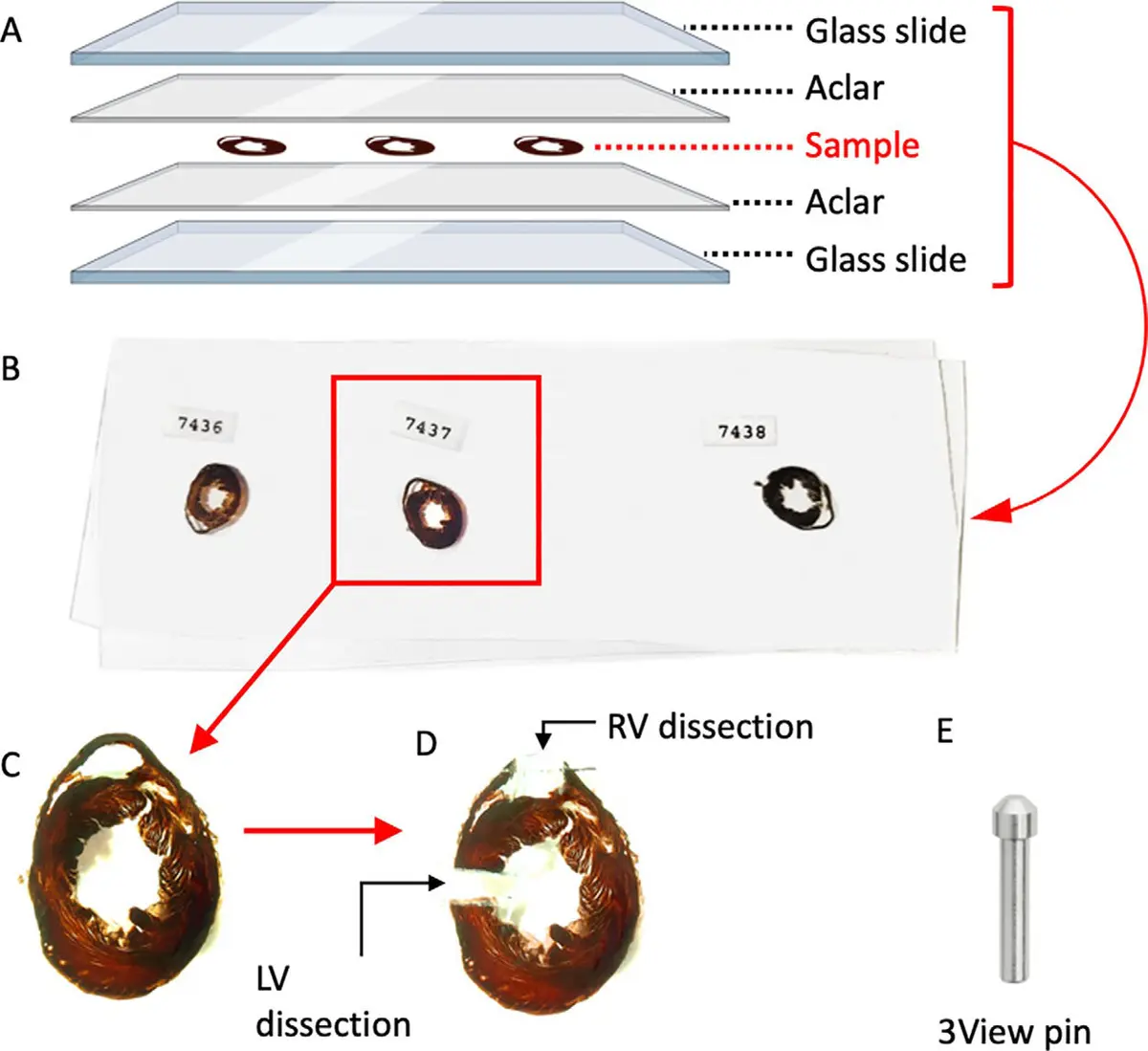
Serial block-face scanning electron microscopy (SBF-SEM) has emerged as an ideal technique to bridge this gap, enabling the automated acquisition of highly aligned, high-resolution 3D data across large tissue volumes spanning hundreds of micrometers [12, 13]. By mounting a miniature ultramicrotome directly within the vacuum chamber of a scanning electron microscope (SEM), researchers can sequentially slice an embedded, heavy-metal stained sample and image the freshly exposed block face using backscattered electron (BSE) detection [17, 32, 34, 161]. This approach circumvents the manual alignment issues of ssTEM and permits rapid, automated data collection [12, 13]. When coupled with optimized *en bloc* staining protocols—such as variants of the osmium-thiocarbohydrazide-osmium (OTO) method—SBF-SEM produces TEM-like image contrast, allowing developmental biologists to trace complex cellular topologies and cell-matrix interactions throughout embryogenesis [12, 32, 33] (Figure 54).
Oogenesis and the Architecture of the Germline Cyst
The foundation of embryogenesis begins with oogenesis, a dynamic process in which mitotic precursor cells, or oogonia, divide and differentiate [161]. In both invertebrates and vertebrates, early oocytes develop within a cellular organization known as the germline cyst, where germ cells remain tightly clustered, interconnected, and enveloped by somatic cells [161]. SBF-SEM has proven exceptionally powerful for investigating the spatial organization of these cysts, particularly in the zebrafish (*Danio rerio*) ovary, where the thickness of the tissue previously limited deep ultrastructural imaging [161].
By acquiring 3D datasets at an ultrastructural resolution, SBF-SEM allows researchers to bypass the limitations of two-dimensional imaging to fully characterize the morphology and connectivity of cyst cells [161]. Three-dimensional volume reconstructions from SBF-SEM data have been used to precisely annotate and visualize cell membranes, centrosomes, and zygotene cilia extending through the cyst [161]. Importantly, this technique has captured critical symmetry-breaking events during oocyte polarization dynamics [161]. For example, prior to symmetry breaking at the leptotene stage, mitochondria are randomly distributed within the cytoplasm; however, at the zygotene stage, SBF-SEM segmentation reveals the explicit polarization of mitochondria aggregating adjacent to the oocyte centrosome [161].
Furthermore, SBF-SEM has shed light on intercellular communication within the cyst. In a mechanism akin to the "dumping" process in *Drosophila*, where nurse cells transfer material to the oocyte via ring canals, SBF-SEM of the zebrafish cyst has identified cytoplasmic bridges (CBs) connecting the cells [161]. Reconstructions of sequential block-face images confirmed the existence of vesicle-like material extending through and within the vicinity of the CB openings, suggesting the active transfer of developmental material between cyst cells [161]. By documenting the number and morphology of these CBs per oocyte, SBF-SEM has provided the first ultrastructural evidence that the zebrafish germline cyst operates as a complex branched network [161].
Early Embryogenesis in Basal Chordates and Marine Invertebrates
SBF-SEM is uniquely suited for the study of whole early-stage embryos and small marine invertebrate larvae, where immense structural complexity is packed into a microscopic volume [256, 258]. The larvacean *Oikopleura dioica*, for example, undergoes rapid larval development characterized by dramatic trunk morphogenesis [256]. Using SBF-SEM, researchers successfully reconstructed the entire trunk region of a hatched tadpole-shaped larva (3 hours post-fertilization) and a young juvenile (10 hours post-fertilization) at the single-cell level [256]. Generating thousands of serial sections, the imaging revealed that while the early larval trunk appeared as an unidentifiable cell mass under a light microscope, SBF-SEM could definitively map the composition and 3D boundaries of the epidermis, brain, inner cells, tail ganglion, and endodermal strand [256].
Similarly, SBF-SEM has resolved long-standing developmental controversies in the cephalochordate amphioxus (*Branchiostoma floridae*) [255, 258]. By analyzing fixed 12-to-13-hour neurula and tailbud embryos, SBF-SEM generated continuous 3D tissue surfaces that mapped the precise distribution of mesodermal, ectodermal, and endodermal cells along the anterior-posterior axis [255, 258]. Crucially, SBF-SEM reconstructions tracked the specific fate of compact mesoderm cells at the posterior extremity of the first left somite [258]. The more dorsal of these cells were shown to give rise to the initial kidney (Hatschek’s nephridium) and participate in branchial nephrogenesis, while the ventral cells became interposed between the ectoderm and endoderm to facilitate the penetration of the larval mouth [258]. Furthermore, by imaging these intact early embryonic tissues, SBF-SEM proved that Hatschek’s right and left gut diverticula—structures widely accepted in classical embryology literature—do not actually form during early development, fundamentally altering proposed homologies of deuterostome head cavities [255]. The technique has also been applied to map the development of the amphioxus notochord [260], the peripheral sensory nervous system in the amphioxus tail [200], and the complete central nervous system connectome of the chordate sibling *Ciona intestinalis* tadpole larva [263].
Embryonic Tissue Ultrastructure and the Extracellular Matrix
During the latter half of embryonic development, cells differentiate and secrete massive amounts of self-derived extracellular matrix (ECM) that extends far beyond the boundaries of individual cells [12]. The ECM acts as a structural scaffold that resists forces of compression and tension, while also providing critical chemical cues and signaling resources for continued cell survival and morphogenesis [12]. Because the mature ECM consists largely of cross-linked, insoluble macromolecules with very slow turnover rates (e.g., collagens in tendon), studying the 3D organization and formation of these dense tissues *in vivo* is refractory to conventional molecular and biochemical techniques [12].
SBF-SEM has drastically improved our knowledge of ECM organogenesis by providing sufficient resolution to trace the geometric parameters of matrices extending over large 3D fields [12]. In the study of embryonic vertebrate tendon development, SBF-SEM allows researchers to track individual collagen fibrils (30–40 nm in diameter) over considerable distances within the developing tissue bundle [12]. For instance, imaging volumes of 14-day embryonic chick tendon and embryonic day 17.5 (E17.5) mouse tail tendon demonstrated that collagen fibrils do not remain strictly columnar, but rather intertwine intricately, mediating highly specific cell-matrix interactions necessary for proper force transmission [12]. Furthermore, SBF-SEM has been utilized to evaluate the 3D tenocyte network in newly hatched chick Achilles tendons [257], to trace the ultrastructural differentiation of Sertoli cells in the developing mouse testis [94], and to characterize the development of the sarcoplasmic reticulum and mitochondrial networks within fetal guinea pig skeletal muscle [35].
Vascular and Renal Organogenesis
The development of the circulatory and excretory systems requires exquisite spatiotemporal coordination, characterized by the fusion of migrating cells and the elaboration of complex membrane architectures. SBF-SEM is increasingly applied to capture these dynamic, transient developmental events [92].
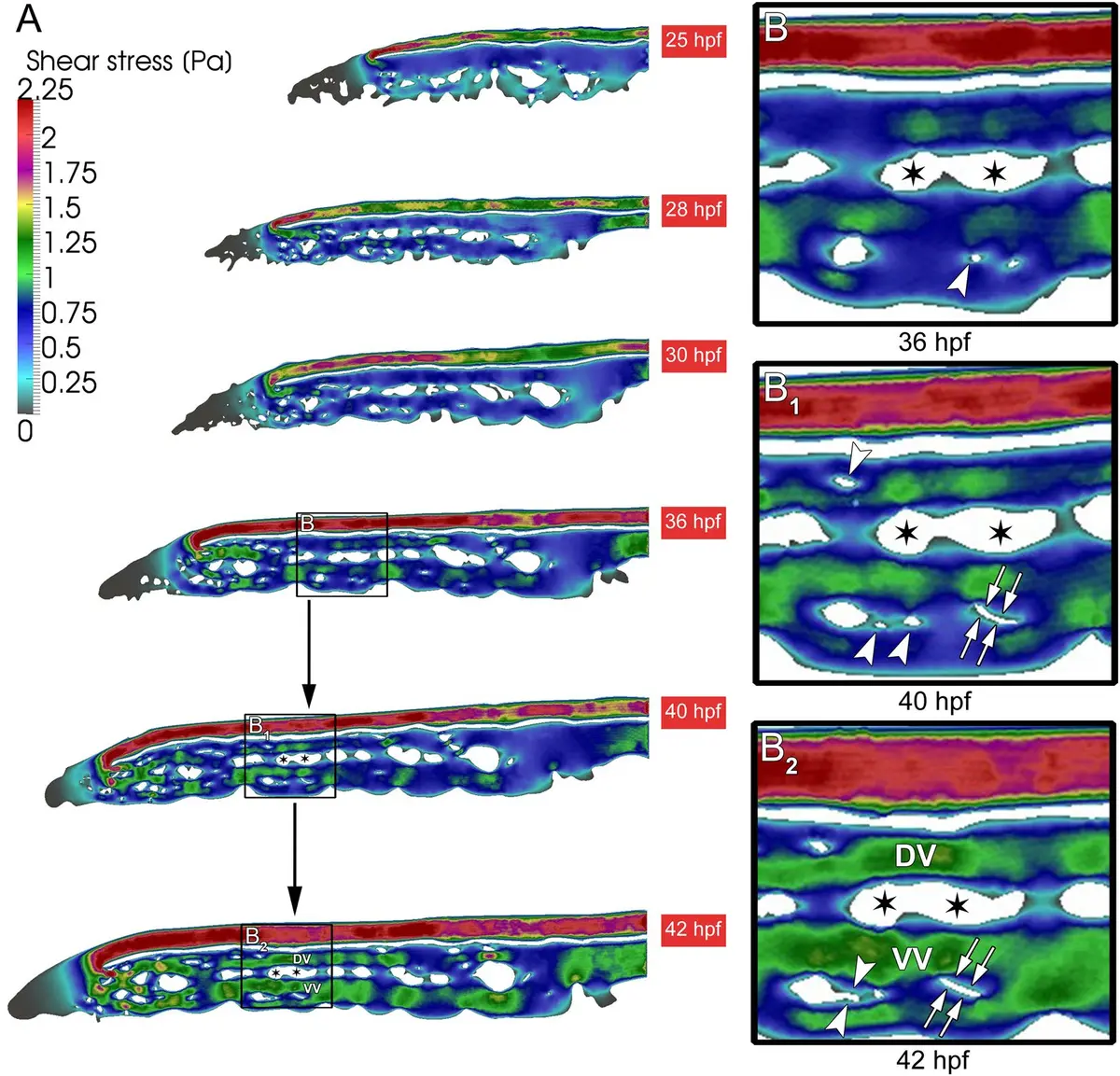
During vertebrate angiogenesis, endothelial cells (ECs) extend long, sensing filopodia toward morphogen gradients [92]. The critical step of anastomosis—where filopodia from two migrating ECs make contact and fuse to form a lumenized blood vessel—is a transient event occurring in an exceptionally small volume (e.g., ~3 ) [92]. Finding such a minute structural event in a large developing tissue is virtually impossible with standard TEM. However, by combining live *in vivo* confocal microscopy of fluorescently labeled ECs in transgenic zebrafish embryos (*Tg(fli1:EGFP)y1*) with volume EM (a correlative light and volume electron microscopy approach), researchers successfully located and reconstructed the 3D ultrastructure of this transient blood vessel fusion [14, 46, 92]. Similar approaches have been employed to study the remodeling of the zebrafish caudal vein plexus, allowing for the 3D reconstruction of intussusceptive pillars and the synergistic interactions between sprouting and intussusceptive angiogenesis during vascular development [259] (Figure 55).
SBF-SEM has also been instrumental in defining the ultrastructure of developing hematopoietic stem cell (HSC) niches. In zebrafish larvae, correlative light and SBF-SEM workflows have localized single, rare HSCs and mapped their structural relationships to surrounding features like pronephric tubules, the glomerulus, and somite boundaries [186]. In the mammalian kidney, the differentiation of podocytes—highly specialized epithelial cells that wrap around the glomerular capillaries to form the filtration barrier—involves massive morphological rearrangements. SBF-SEM imaging of neonatal rat kidneys has successfully captured the developmental progression of podocytes, mapping the whole-cell architecture and the 3D elaboration of their complex foot processes in an automated, high-throughput manner previously impossible with conventional SEM or TEM [40, 68].
Developmental Connectomics and Neurogenesis
One of the most prolific applications of SBF-SEM is in the field of connectomics and developmental neurobiology [12, 13, 32, 42]. Because SBF-SEM routinely achieves a lateral resolution of 5–20 nm and a typical section thickness of 25–50 nm, it is capable of resolving the thinnest unmyelinated axons, dendritic spines, and individual synaptic vesicles [13, 32].
In studying the assembly of neural circuits during development, volume EM provides the 3D context needed to identify how physical connections between neurons are established and refined [184, 188]. For example, SBF-SEM has been used to elucidate neural circuit assembly in the developing mouse retina, utilizing skeleton tracing and 3D surface reconstructions to identify precisely where synaptic sites form among maturing retinal cells [188]. In the central nervous system, researchers have mapped the postnatal development of cerebellar granule neurons, tracking their morphological maturation and the formation of dendritic claws [262]. Other studies have monitored migrating neuroblasts in the postnatal brain, using SBF-SEM to capture the dynamic changes in the ultrastructure and orientation of the primary cilium during different phases of cell migration [8, 155].
Beyond mere neuronal wiring, SBF-SEM is revealing novel interactions between the developing nervous system and surrounding non-neural support cells. High-resolution 3D ultrastructural analyses of the embryonic (E12.5 and E14.5) mouse neocortex have uncovered surprising interactions between the developing vasculature and neurogenesis [207]. SBF-SEM reconstructions revealed that endothelial tip cells projecting from the vasculature extend long, slender processes that physically traverse neighboring pericytes and penetrate deep into apical and basal neural progenitor cells [207]. Such invasive cell-cell interactions during neocortical development highlight communication pathways that would be indistinguishable without the volumetric context provided by SBF-SEM [207].
Additionally, SBF-SEM has contributed heavily to mapping the developmental timeline of simpler model organisms. By acquiring serial section and SBF-SEM datasets of the *C. elegans* brain at various developmental stages, researchers have generated wiring diagrams that demonstrate how the complexity and relative strength of synaptic connections evolve as the animal matures, even while the overall physical geometry and neuron positions remain relatively constant [9]. The technique has similarly been used to study myelin biogenesis and the resolution of pathological ultrastructure by microglia in developing zebrafish [132], and to map the complete morphological development of TmA visual neurons in the optic lobes of fourth instar locusts [261].
Conclusion
By filling the critical gap between optical microscopy and conventional transmission electron microscopy, Serial Block-Face Scanning Electron Microscopy has triggered a paradigm shift in developmental biology [12, 14]. Whether revealing the spatial dynamics of the germline cyst, the 3D architecture of the embryonic extracellular matrix, the fusion of developing blood vessels, or the highly complex wiring of the maturing nervous system, SBF-SEM allows researchers to reliably correlate form and function across massive tissue landscapes [92, 161, 188]. As data analysis pipelines and automated segmentation algorithms continue to improve [100, 149], SBF-SEM will undoubtedly remain a cornerstone technique for investigating the ultrastructural mechanics of organogenesis and embryology.